Benchling Product Bulletin: Spring 2021
Benchling’s customers are leading innovation in many transformative areas of science, which promise to improve human health and environmental sustainability. At Benchling, one of our top priorities is to support new and emerging modalities through our Life Sciences R&D Cloud. As a recent example, in the spring release we introduced functionality for modeling RNA therapeutics.
RNA has emerged as a promising approach for addressing biological targets previously thought to be “undruggable.” A key factor driving R&D in this space is RNA oligonucleotides (oligos). These short, synthetic strands of RNA support a wide range of applications, from guiding gene editing in CRISPR to detecting nucleotides during in-situ hybridization.
Benchling supports scientists with tools for conducting experiments on the cutting edges of science, including this powerful new modality. Scientists working in Benchling can now design and bind RNA oligos to complex biomolecules, and Benchling will continue to offer new capabilities for chemically-modified RNA therapeutics.
RNA oligos now supported in Benchling
The new RNA oligo capabilities within Benchling’s Molecular Biology application makes modeling therapies like siRNA and CRISPR sgRNA more efficient, and simpler to integrate into your workflow. With a centralized data model created specifically for these complex molecules, this functionality makes scientific results and insights easier to access and contextualize.
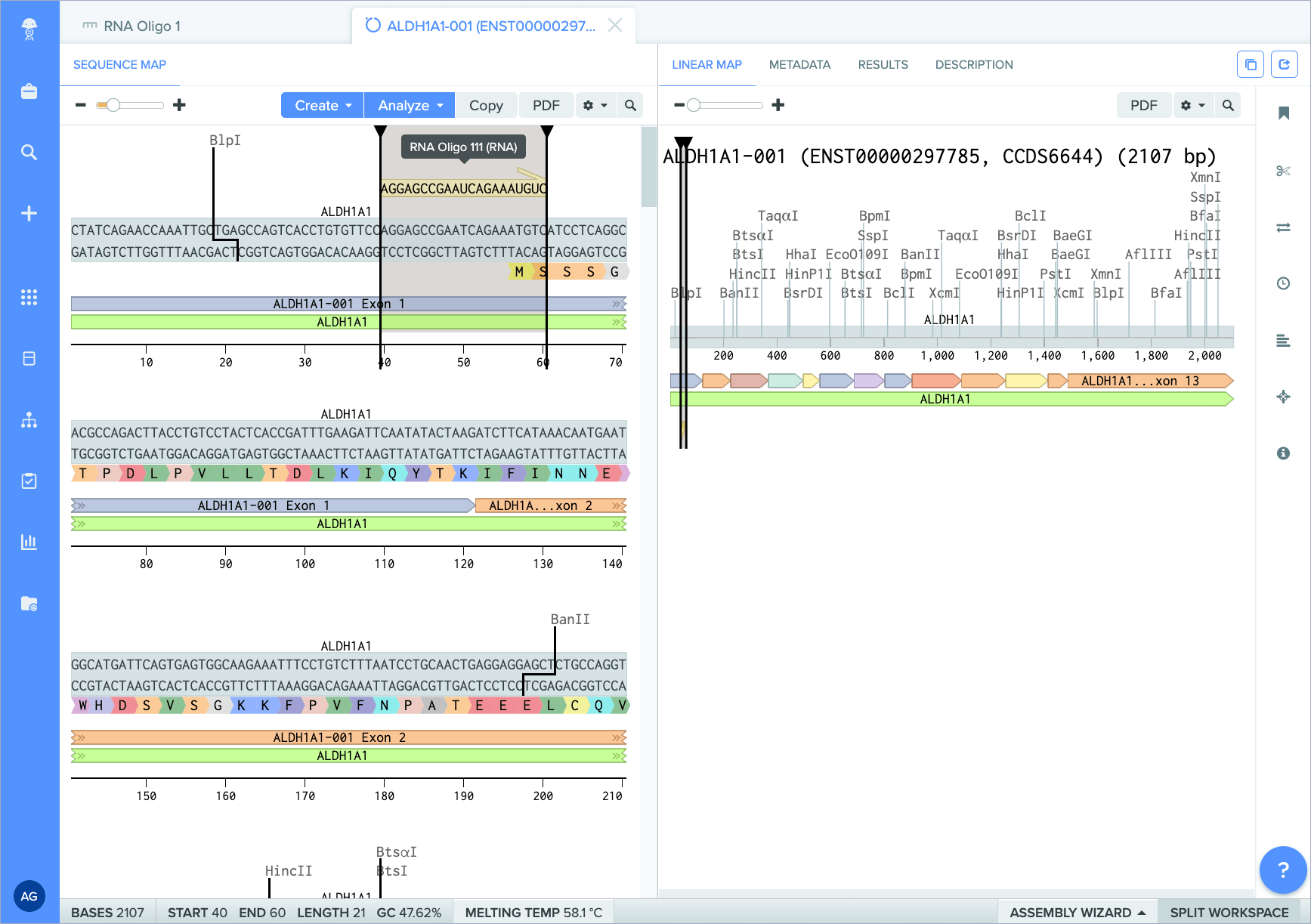
How this feature helps you
Previously, modeling RNA oligos required time-consuming workarounds, such as using DNA oligos, custom entities, or even siloed software solutions. Now, the new RNA oligo data type in Benchling streamlines experiment design, while upholding the integrity and accuracy of your data across all Benchling applications.
Key functionality
This comprehensive tooling for RNA oligo R&D enables you to:
Create and register new RNA oligos from global create, Registry, Notebook, or the CRISPR guide tool
Model RNA-based oligo therapies with RNA-specific design tools and data fields
Treat RNA oligos as primers, and attach them to DNA sequences for modification
Leverage oligo-specific endpoints for DNA and RNA, replacing the previous generic oligo endpoint
Use cases for RNA oligos
With RNA oligos, scientists can keep pace with the latest innovations in biotech, such as:
Modeling RNA oligo-based therapies
Leveraging RNA oligos as probes for in-situ hybridization
Using RNA oligo tools to design sgRNA, for use in CRISPR gene editing
Notebook
Benchling Notebook’s easy-to-use interface lets you log experiments, track information, and capture sample results, all while driving unified data capture across Benchling. In this release, we’ve improved Notebook tables and added a more user-friendly interface for editing table rows and modeling chemical structures.
Add multiple rows to structured tables
Users can now add multiple rows to structured tables by double clicking on the “drag to add” bar. This makes it easier and faster to add large numbers of new rows at once.
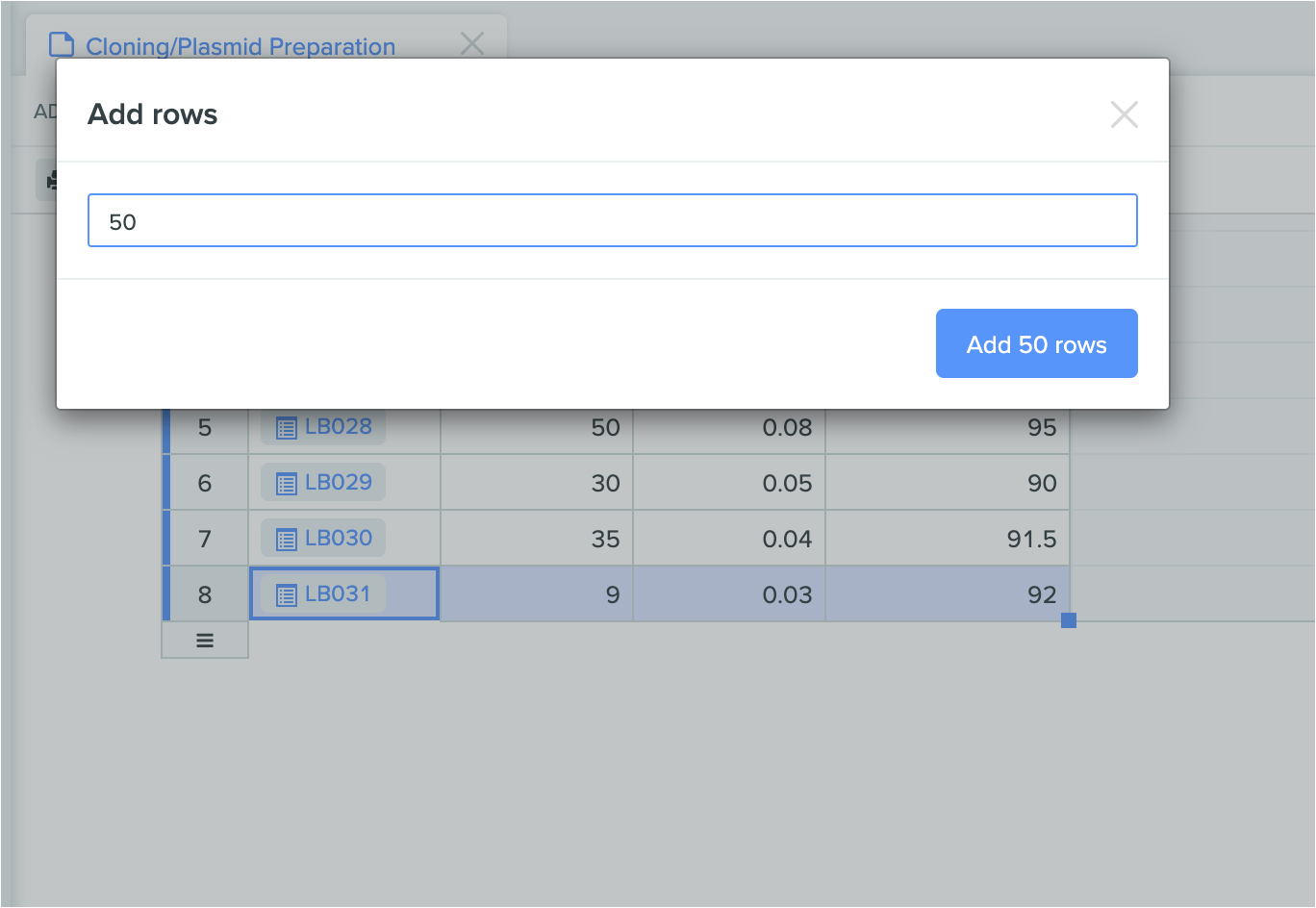
New ways to model chemical structures
MolFiles in Notebook entries: you can now copy compounds created in drawing tools like MarvinDraw and ChemDraw, and paste the corresponding MolFile text directly into an entry. Notebook will automatically detect this file and will initiate an upload of the compound’s image.
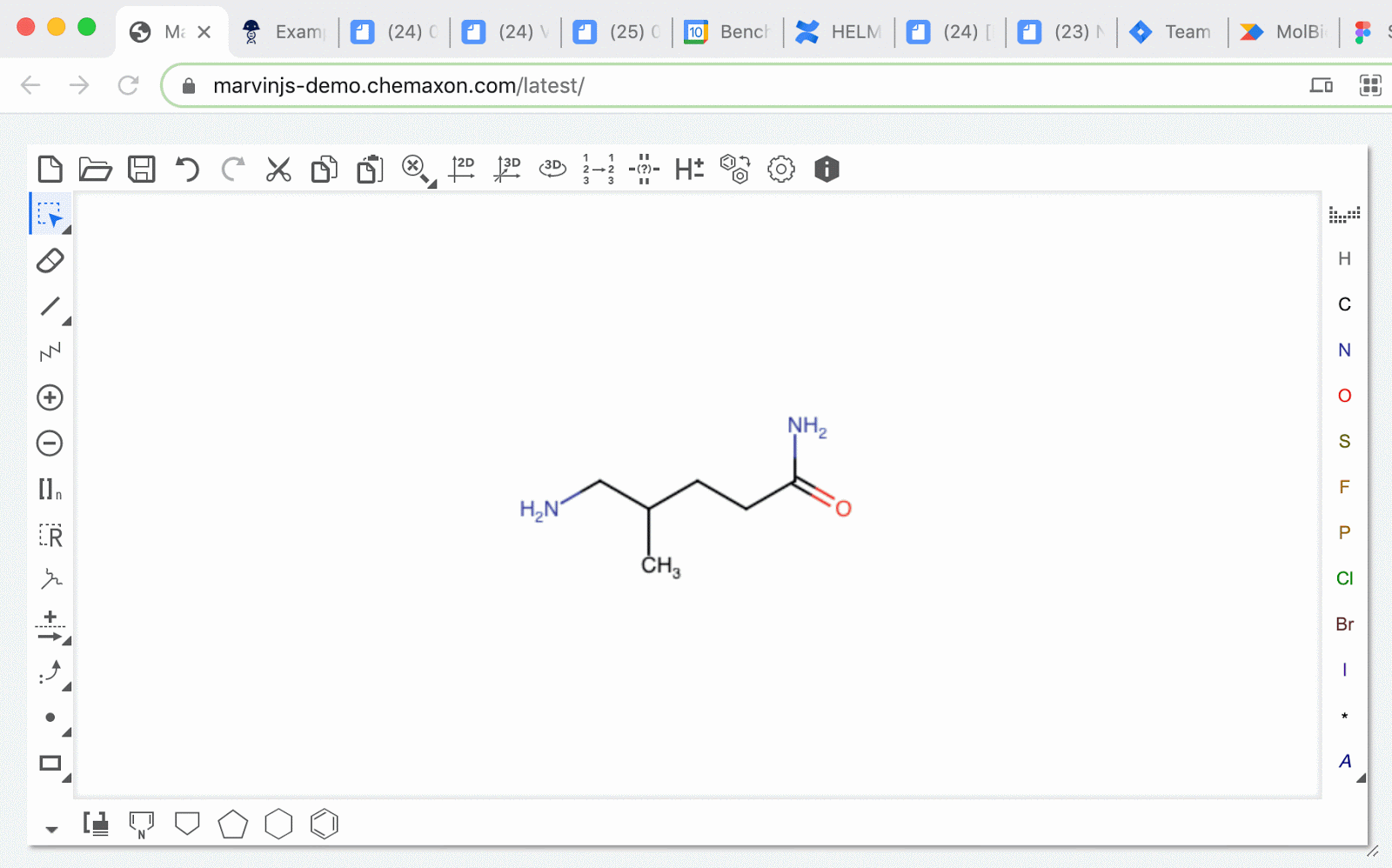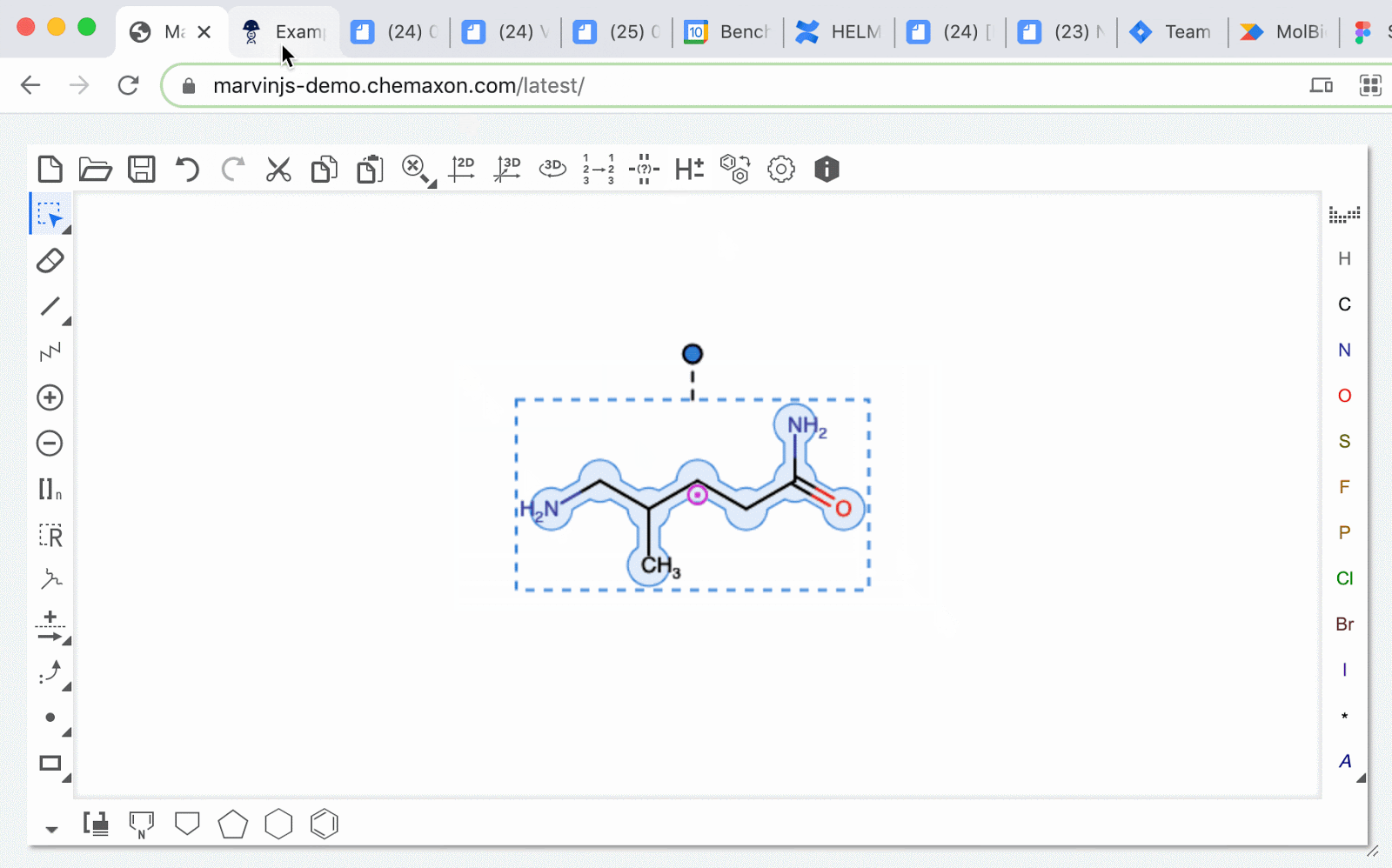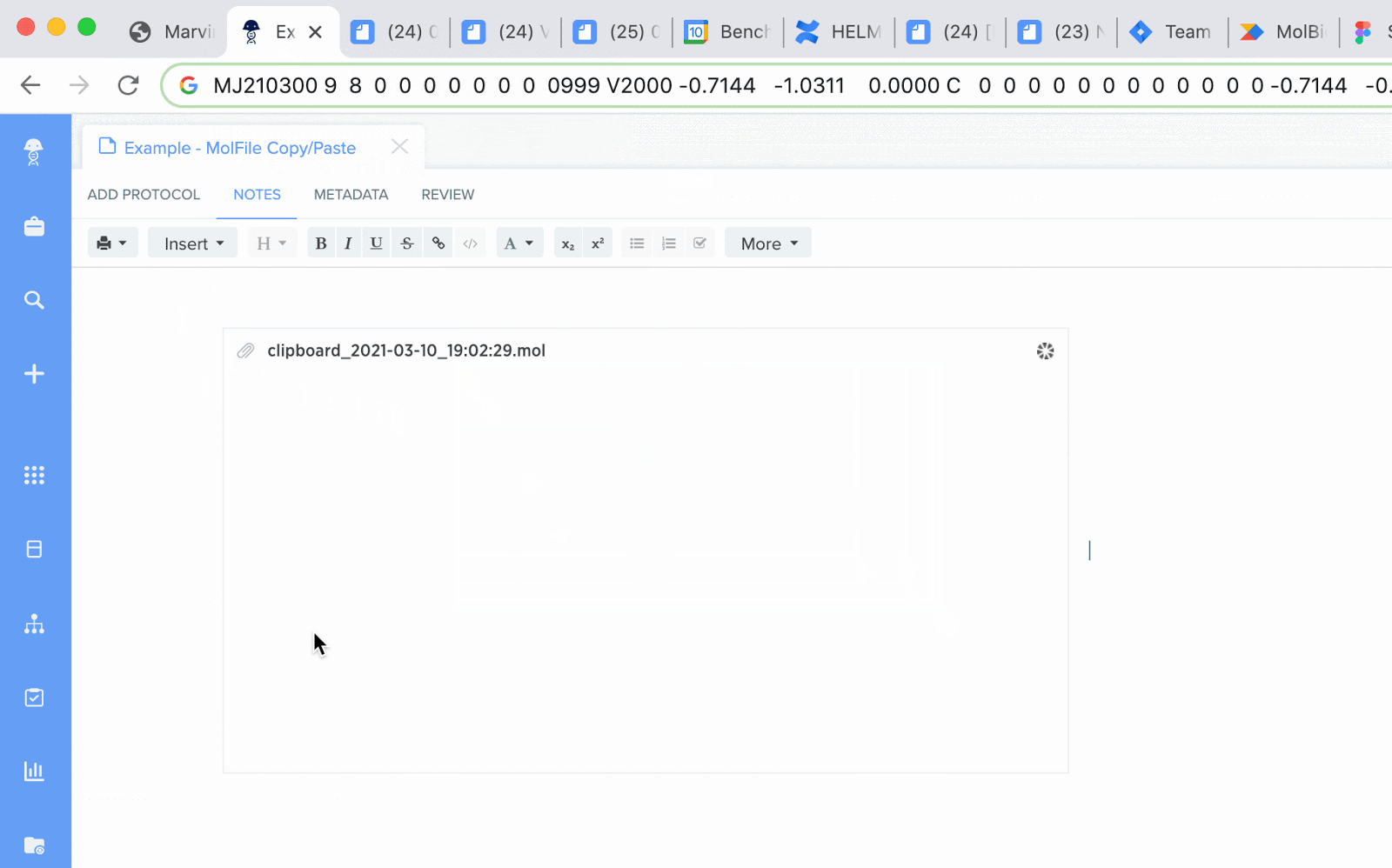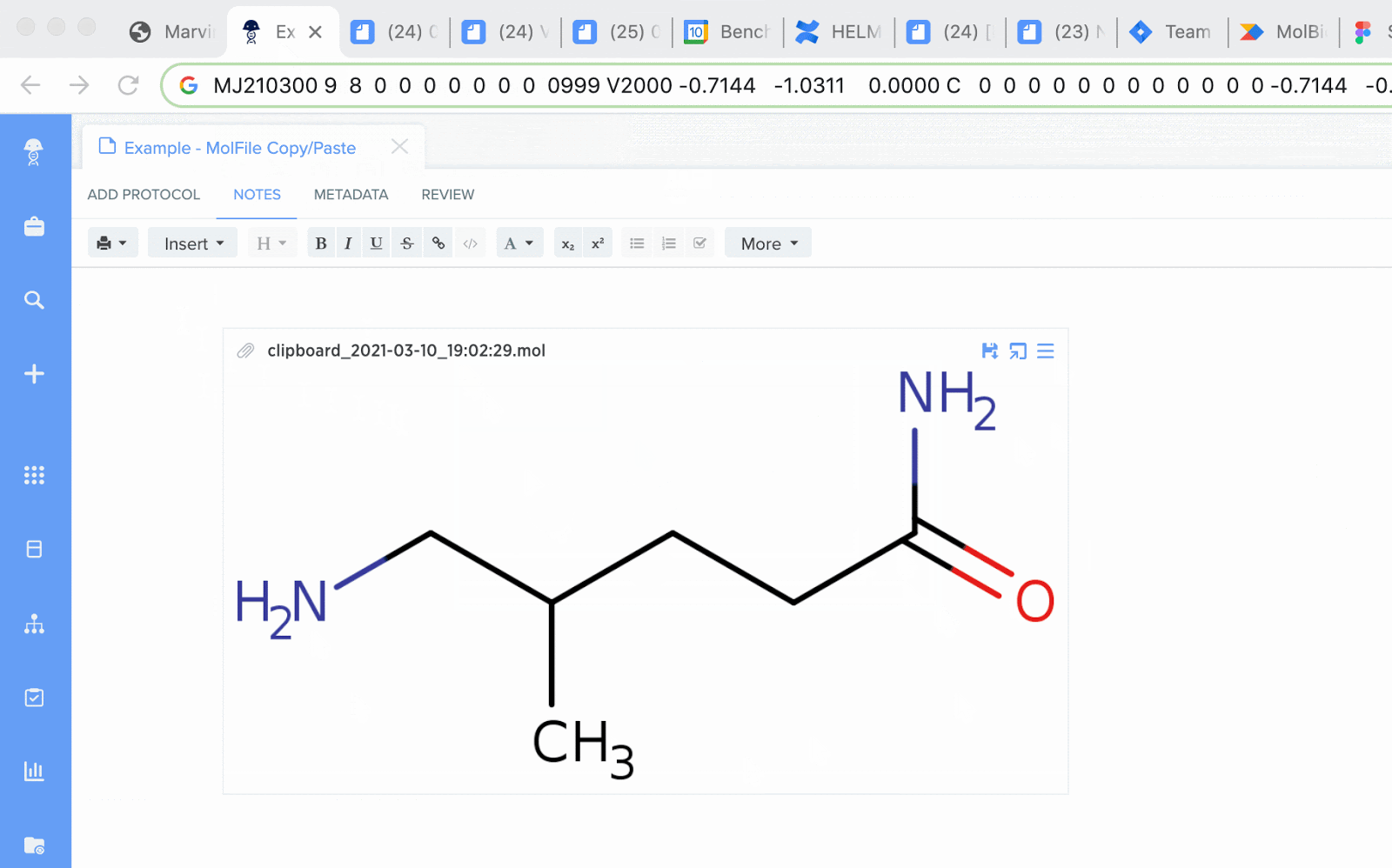
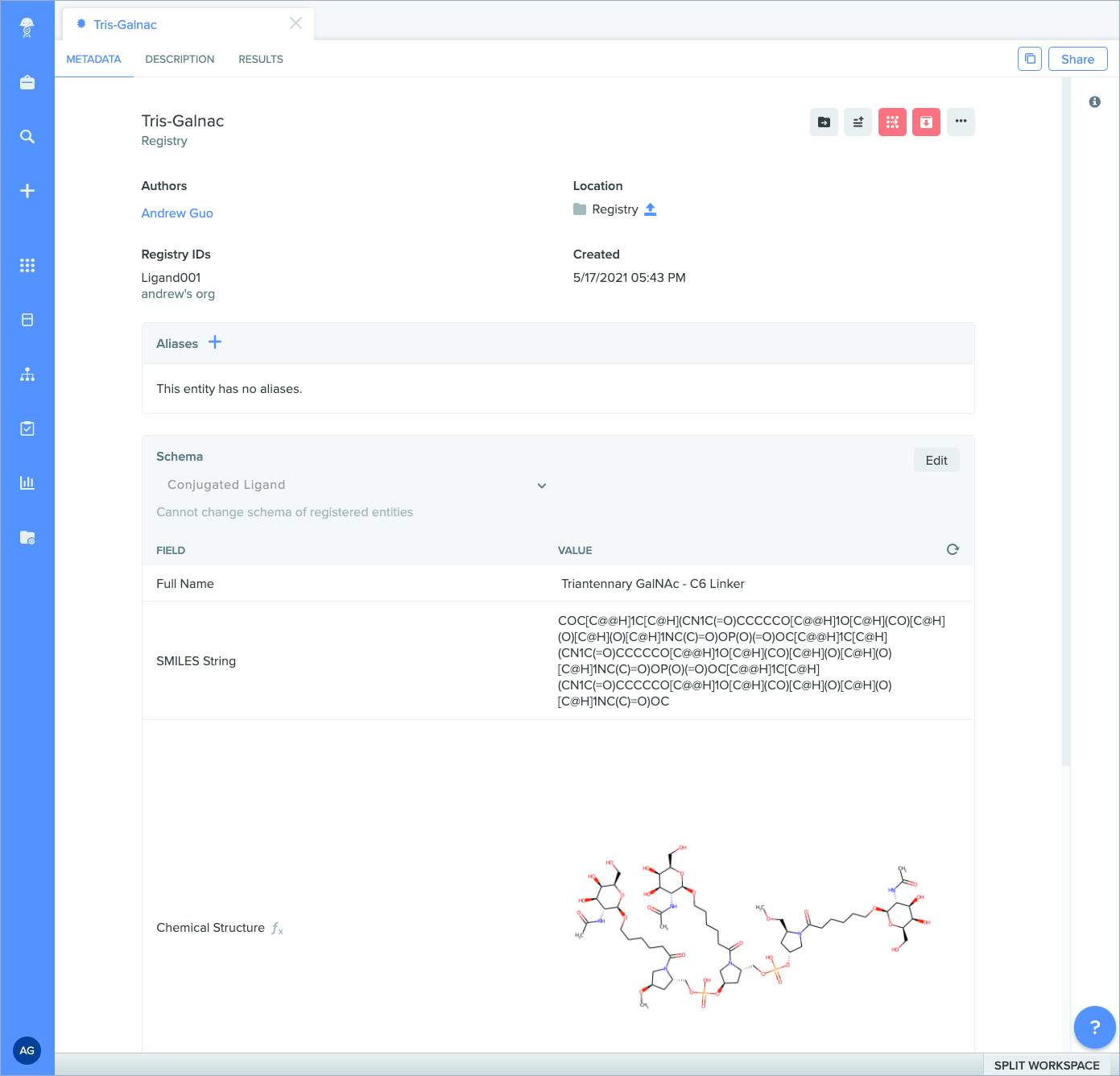
SMILES strings as chemical structure: schema fields containing SMILES strings can now be rendered as an image preview of the chemical structure. The newly generated image will appear in the corresponding blob field.
Molecular Biology
Benchling Molecular Biology is an in-silico design tool that supports a wide array of molecular modeling workflows, such as primers, digests, and CRISPR guides. In addition to RNA Oligos mentioned above, we’ve simplified how you update oligo information during data import.
Import/export enhancements
You can now maintain sequence information in one place, and import entity data more quickly and easily, with the following features:
GenBank annotations: Any DNA sequence that you export as a .gb (GenBank) file will now contain GenBank annotations, which reference specific sequence components and translations.
Bulk entity updates: When importing new entity data via spreadsheet or CSV, you can now update DNA sequences and DNA/RNA oligos in bulk.
Registry and Inventory
Benchling Registry and Inventory work together to power your experiments through connected data — whether that’s data on sequences, samples, or experimental results. With Benchling Registry, you can standardize, connect, model, and track samples across your experiments. Benchling Inventory provides a digital window into your physical lab, enabling you to track vials, wells, batches and more, while linking each of those elements to relevant results. In this release, we increased the number of supported values in mixture tables, and added new ways to configure mixture experiments.
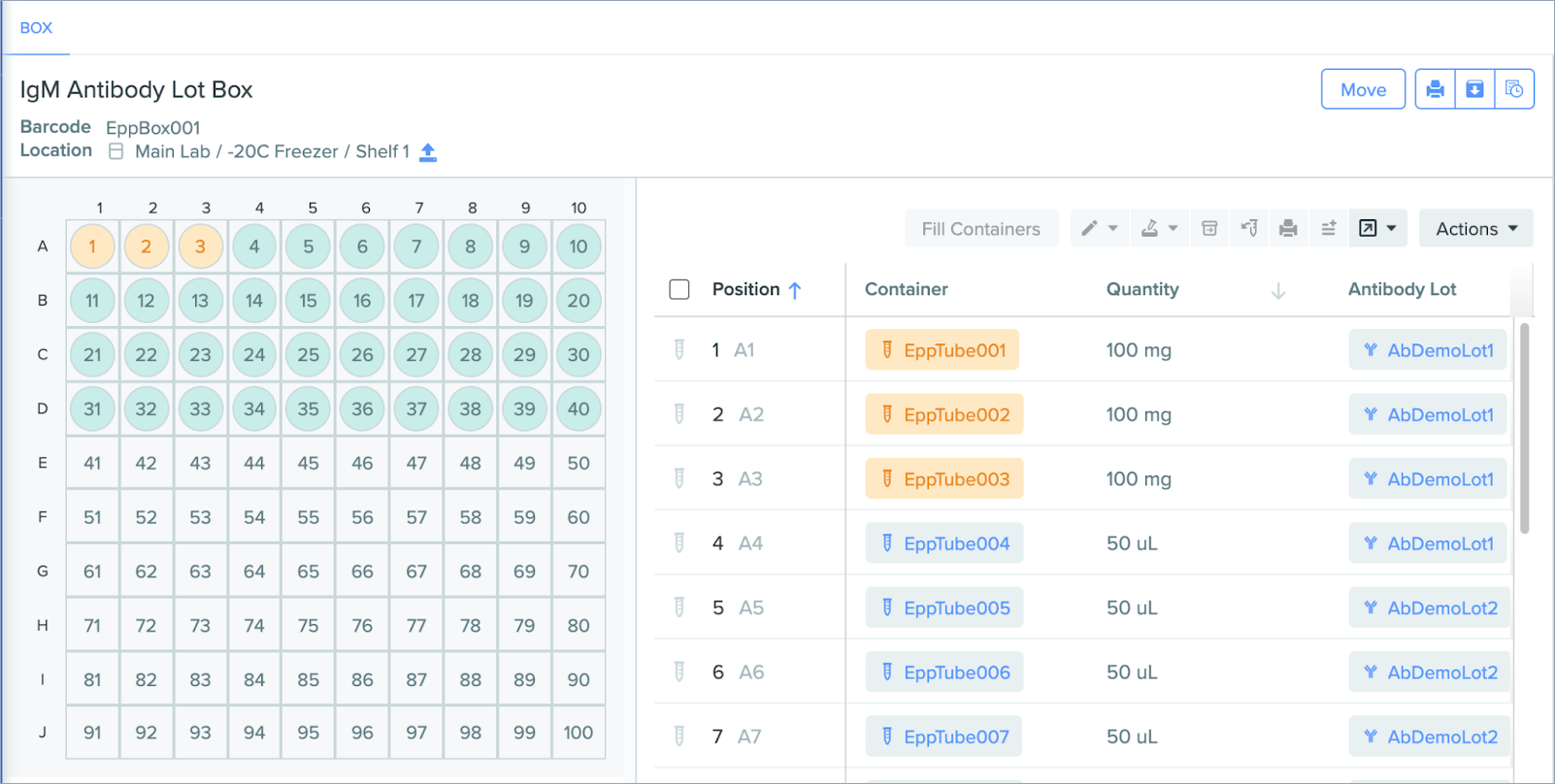
Support for solid units
Container entities now support solids in addition to liquids. This means users can now track solid substances such as powders, seeds, and chemical reagents, making it easier to execute a wider variety of use cases within Benchling. To facilitate this change, the default unit of measurement for containers is now “quantity” instead of “volume.”
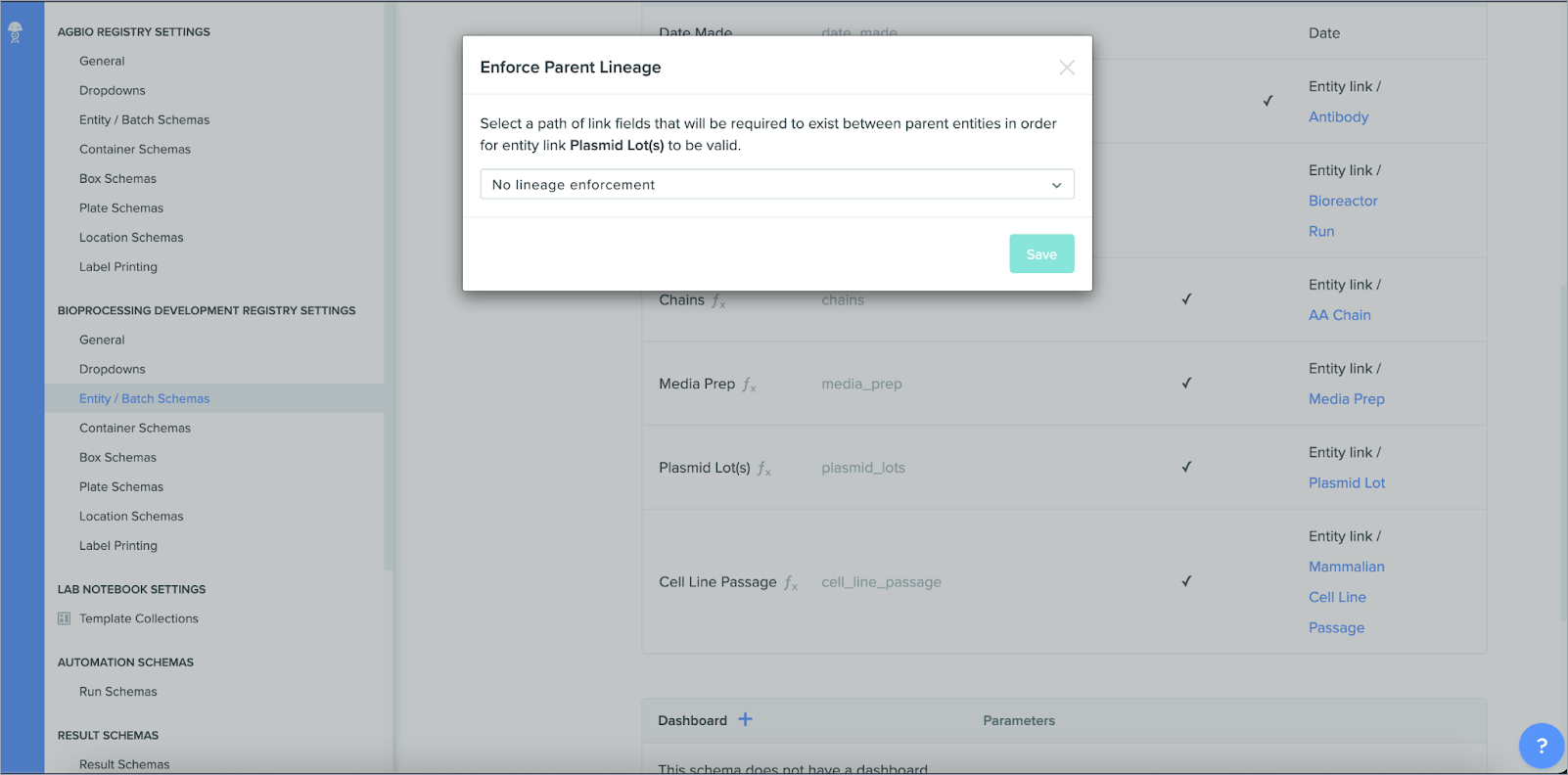
Enforcing entity lineage
You can now enforce relationships between child/parent entities to ensure accurate linkage between items across different areas of Benchling. For example, when modeling a plasmid that results in a specific protein, the system will validate that the plasmid prep links to the correct resulting protein lot. Enforcing lineage between entities helps maintain full traceability and accuracy of experiments and biological data, upholding the integrity of your data while minimizing the potential for human error.
Create Registry IDs and names simultaneously
You can now set registry IDs and names at the same time by creating and deploying a specialized name template. This streamlines the creation of entities, and aligns your team around one unified process for registering and naming new entities.
More visibility into search results
Search results in Registry and Inventory now contain more information about your entities and containers, making it easier to view and access relevant data. Search results now also display:
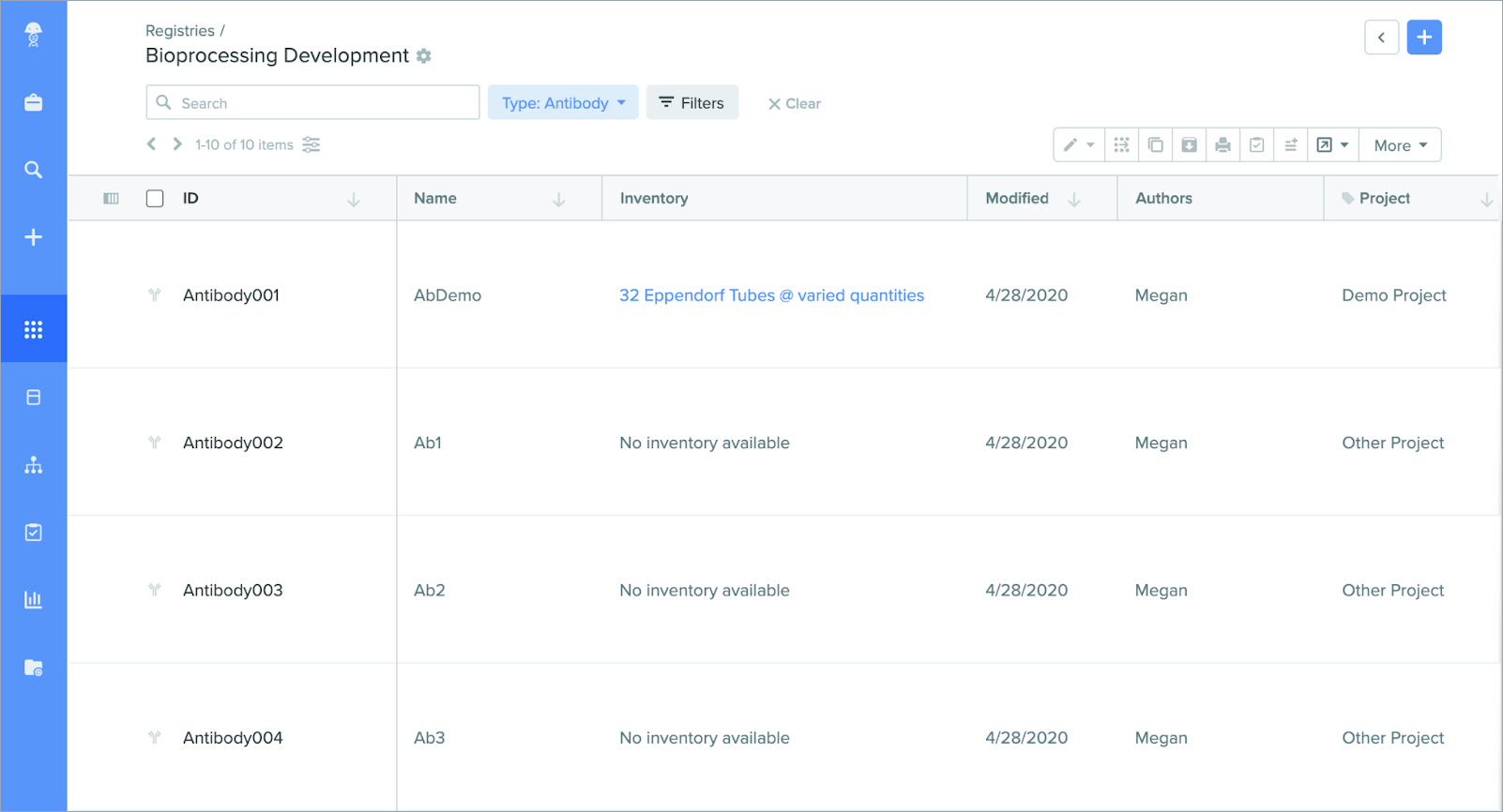
Inventory information: You can now view aggregate Inventory information alongside an entity on the search results page. When you perform an entity search in Registry, a new column in your search results shows available Inventory items for each line. Links in this column will take you directly to a pre-filtered Inventory search, making it easier to navigate to the relevant container entries.
Mixture information: When you search for mixtures, your search results now include more detailed mixture information, such as related amounts and units.
Concentration information: You can now view concentrations for containers with one item through the expanded Inventory search.
New ways to link information across your Benchling data
Inventory now supports entity, entry, and blob links. This lets you seamlessly connect Inventory items to Registry schemas through container, plate, box and location links.
Insights
With Insights, you can visualize, query, and share Benchling data to help generate actionable insights about your R&D pipeline, experiments, team productivity, and resource allocation. In this release, we’ve brought Insights help documentation directly into the application and improved the user interface for dashboards and templates.
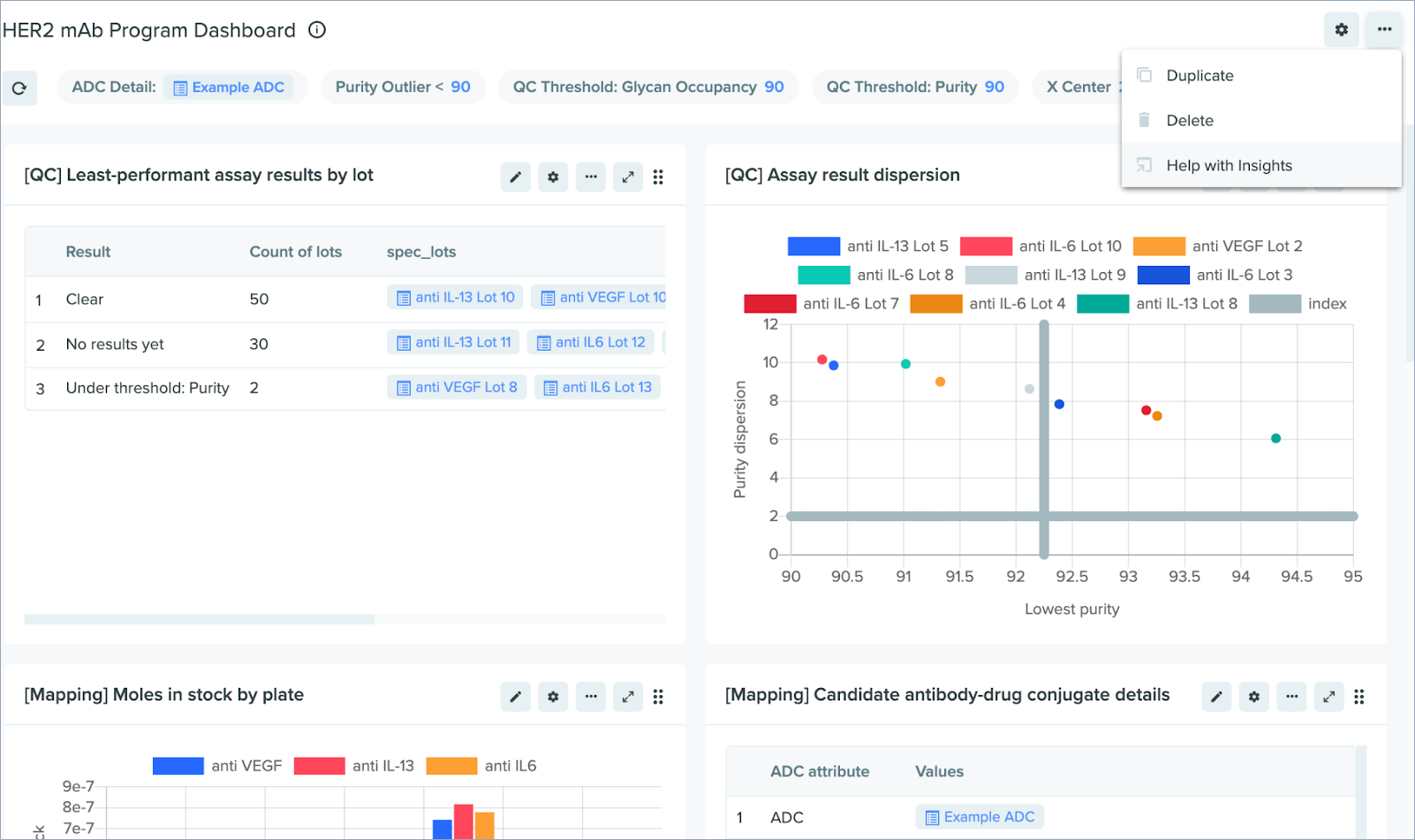
Help documentation in Insights
We’ve added links to help documentation in Insights and made these links available from both the main dashboard menu and the SQL block menu. The help docs provide information on a wide variety of Insights topics: writing SQL, managing permissions, configuring visualizations, and more.
Insights UI enhancements
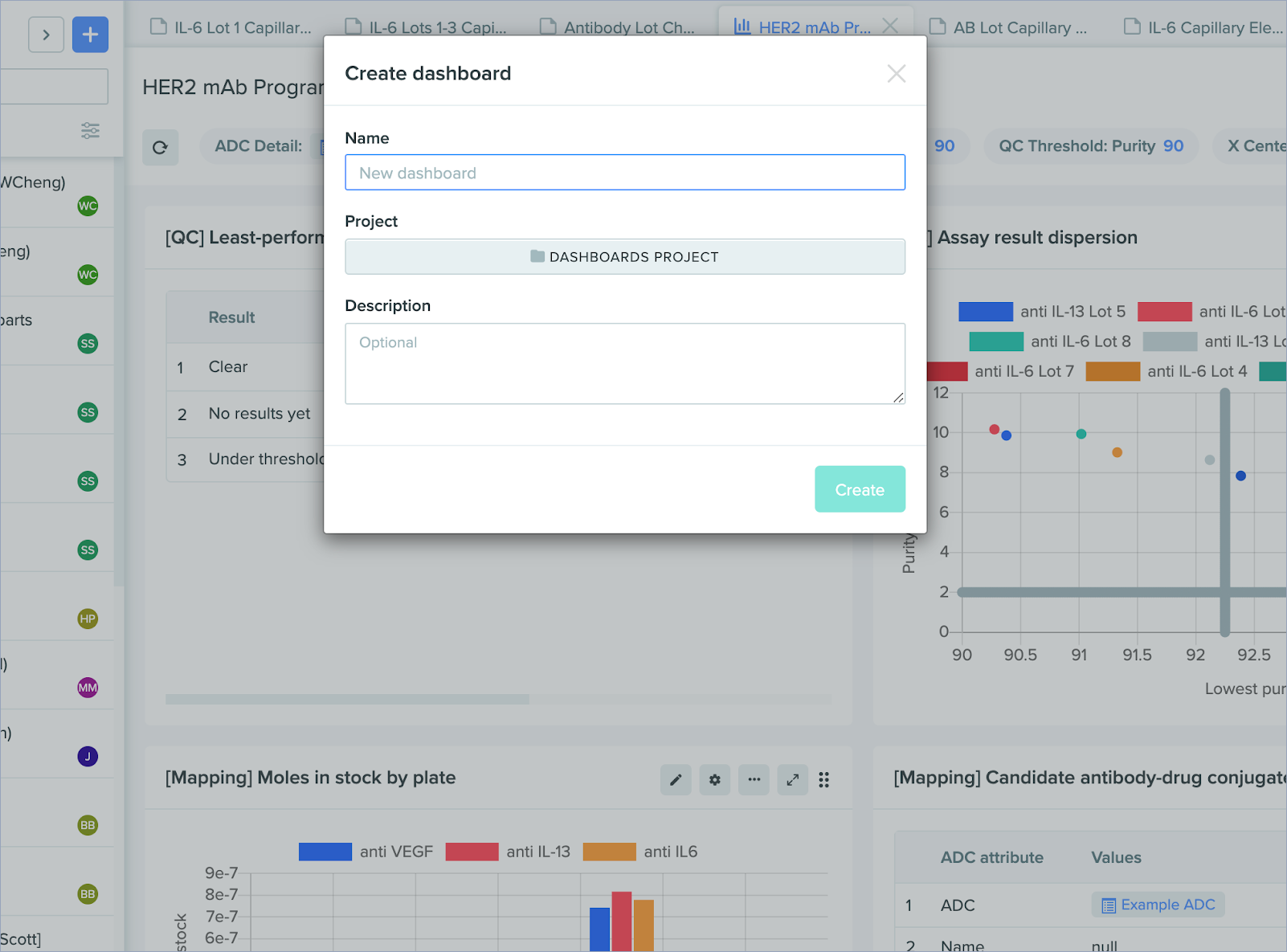
You can now build and edit Insights templates and dashboards faster, with the following UI updates:
Add Insights dashboards from global create: Create new Insights dashboards directly from the global create menu.
Clickable rows in SQL templates: Select, unselect, and group multiple rows in an Insights SQL template.
Lab Automation
Lab Automation lets you integrate lab instruments — like liquid handlers and plate readers — into Benchling to scale experiments, increase scientist productivity, and sync research across software and hardware. In this release, we’ve made it easier to configure runs and keep results data consistent and clean across the Benchling platform.
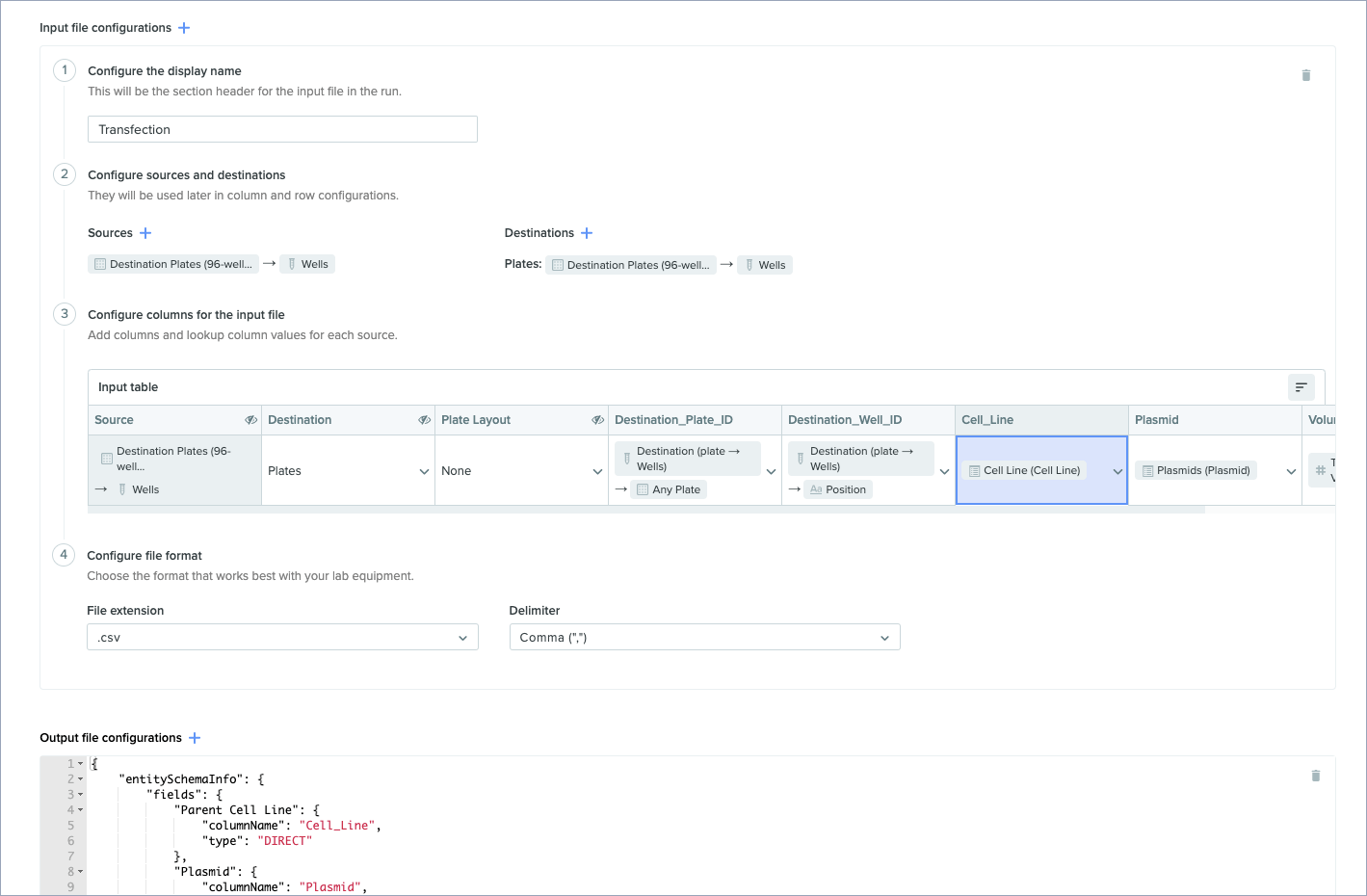
Codeless configuration for Lab Auto runs
Configure Lab Automation runs more quickly and more easily with the Run Schema UI. This updated schema lets you configure run fields and input file config via the user interface, without needing to use JSON. (Please note: the output file config still requires JSON to configure.)
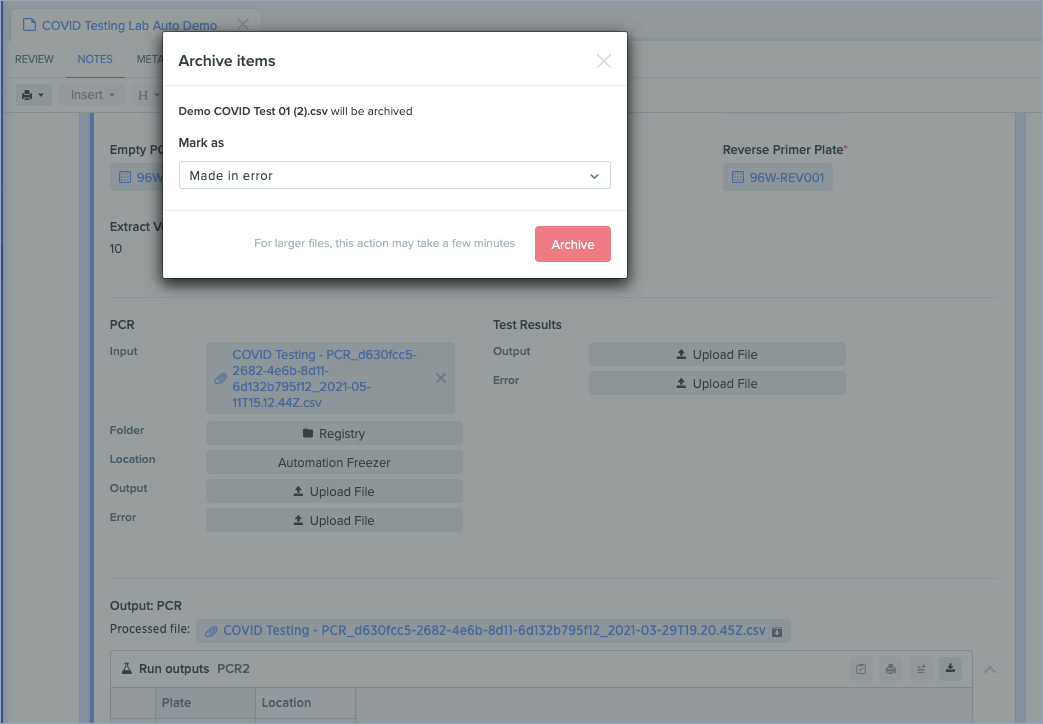
Archiving run results and uploading multiple files
You can now archive results created during Lab Automation runs and upload multiple results files once the previous output configuration file is processed. This makes it easier to manage results data, and helps maintain clean information across the Benchling platform.
New plate layout functions
This release includes two new functions that provide more options for mapping plate layouts.
REVERSE_INTERLEAVE plate layout
: This function performs the reverse of the INTERLEAVE function, allowing users to specify a source plate that is larger than the destination plate. Originally, the REVEARSE_INTERLEAVE plate layout only supported multiples of four. In this release, you can use any size plate layout.
REVERSE_STAMP_TO_QUADRANTS plate layout
: Within a plate layout, this function performs the reverse of the PLATE_STAMP layout. That is, you can now map larger plates to the corresponding locations on smaller plates (for example, stamping a 384-well plate onto a 96-well plate).
More ways to configure your runs and manage results
You can now configure runs more quickly with a streamlined set of processing steps, and use results syncing to facilitate data parity across Benchling.
Support for non-specific schemas
: When using SOURCE_CONTENTS to create new entities, you no longer need to define the schema, as the entity creation template now accepts non-specific schemas.
New find-and-replace functionality
: Quickly find and replace cell contents by using the FIND_REPLACE processing step in Lab Automation run configurations.
Support for blob links in output file processing: You can now insert blob links into Lab Automation results tables, eliminating the need to manually copy and paste this information.
New DUPLICATE processing step: You can copy contents of a specific column in an output configuration file, and configure Benchling to automatically port them over to a new column at the end of the resulting Lab Auto file. This processingStep accepts the following arguments: columnToDuplicate (the name of the column to duplicate) and newColumnName (the name of the new column where the contents will be copied).
Syncing between run_id and results tables: When using warehouse tables in Insights, you can now view run_id and entry_ids associated with your results.
Developer Platform
The Benchling Platform connects capabilities and data across Benchling to provide a simple, unified user experience. In this release, we’ve launched a new-and-improved API documentation page, and have continued to extend the flexibility of the developer platform through updated schema permissions and APIs.
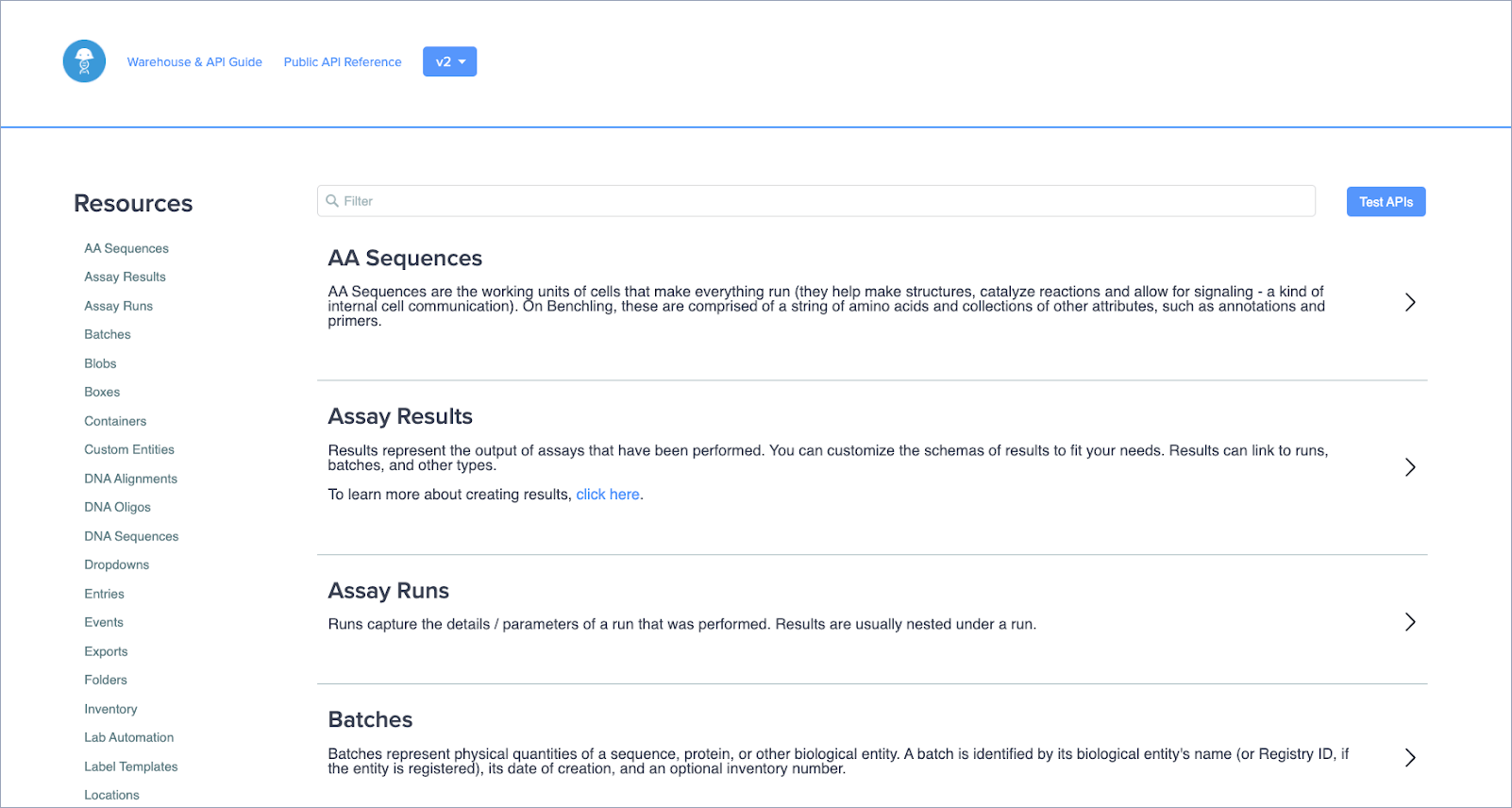
New interactive page for API documentation
Access fully open API documentation that updates in real-time as Benchling APIs evolve, guaranteeing your APIs are always current. You can also make calls directly from the documentation for faster adoption of any Benchling API. View this documentation at benchling.com/api/reference.
More control over users in Benchling
Admins can now manage access to each app within the Benchling platform, providing tighter control over your user privileges. In the tenant admin console, Admins can provision either full access or no access for any user, team or org within Benchling. The ability to manage users more granularly ensures that only authorized employees are able to access each app within your system. Please note: You can toggle access on/off for any Benchling application except the Notebook, which all users must have full access to.
Event support for AWS GovCloud
The Events API functionality triggers external systems from events within the Benchling platform. This allows for a seamless flow of information across your systems by notifying external tools of changes happening in Benchling. In this release, AWS GovCloud can now subscribe to events in Benchling.
API updates
Benchling APIs are built to match the speed and agility of modern R&D, making it easy to extend Benchling into your broader digital ecosystem. We regularly add new APIs to further increase Benchling’s flexibility and expand its range of possible integrations. In this release, we’ve included new ways to filter and interact with endpoints and made it easier to copy API IDs for use in your platform.
Python SDK expansion: You can now use a Python SDK to interact with Registry and Storage endpoints.
Support for “nameIncludes” filter: Listing endpoints that support the “name” filter now also support the “nameIncludes” filter. The new endpoints include AA sequences, DNA sequences and DNA oligos.
“name” filter added to endpoints: Endpoints for folders, label templates, label printers, projects, and registries now support the “name” filter.
Updated URLs for Notebook API: Notebook template URLs now contain the full API ID.
“Copy API ID” button: You can now toggle an on/off a button for copying API IDs. This lets you copy API IDs by right-clicking on the row where they’re listed.
Create Notebook entries with attachments: You can now use the API to create Notebook entries with attachments.
Public API IDs for schema fields: The API IDs for schema fields are now exposed in the public API. This allows integrations and scripts to reference a stable identifier that won’t be thrown off by a name change.
Powering breakthroughs for over 1,200 biotechnology companies, from startups to Fortune 500s
