Benchling Product Bulletin: Fall 2021
As we continue to evolve Benchling’s R&D Cloud, we’re focused on improving how information flows between scientific teams to streamline processes, boost collaboration, and accelerate informed decision making.
In our most recent product update, we released dozens of features that simplify how you share, reference, and use data in Benchling. We’ve updated results tables in Notebook to optimize results capture for entries, plates, and containers. In Insights, teams can quickly visualize powerful trends from these results with new heat maps and IC50 response curves on scatter plots. For teams leveraging Workflows, task groups functionality makes organizing and tracking jobs and their related data easier, bringing deeper process-level intelligence to your work.
Read on for more information about these features, including a spotlight on new lookup tables, which facilitate easier access to key data across Benchling.
The latest Benchling release also delivers a number of helpful features across all of our cloud applications.
Notebook
Benchling Notebook’s easy-to-use interface lets you log experiments and protocols, track information, and record sample results in Benchling's centralized data model. In addition to new lookup tables detailed above, we’ve also reimagined results tables.
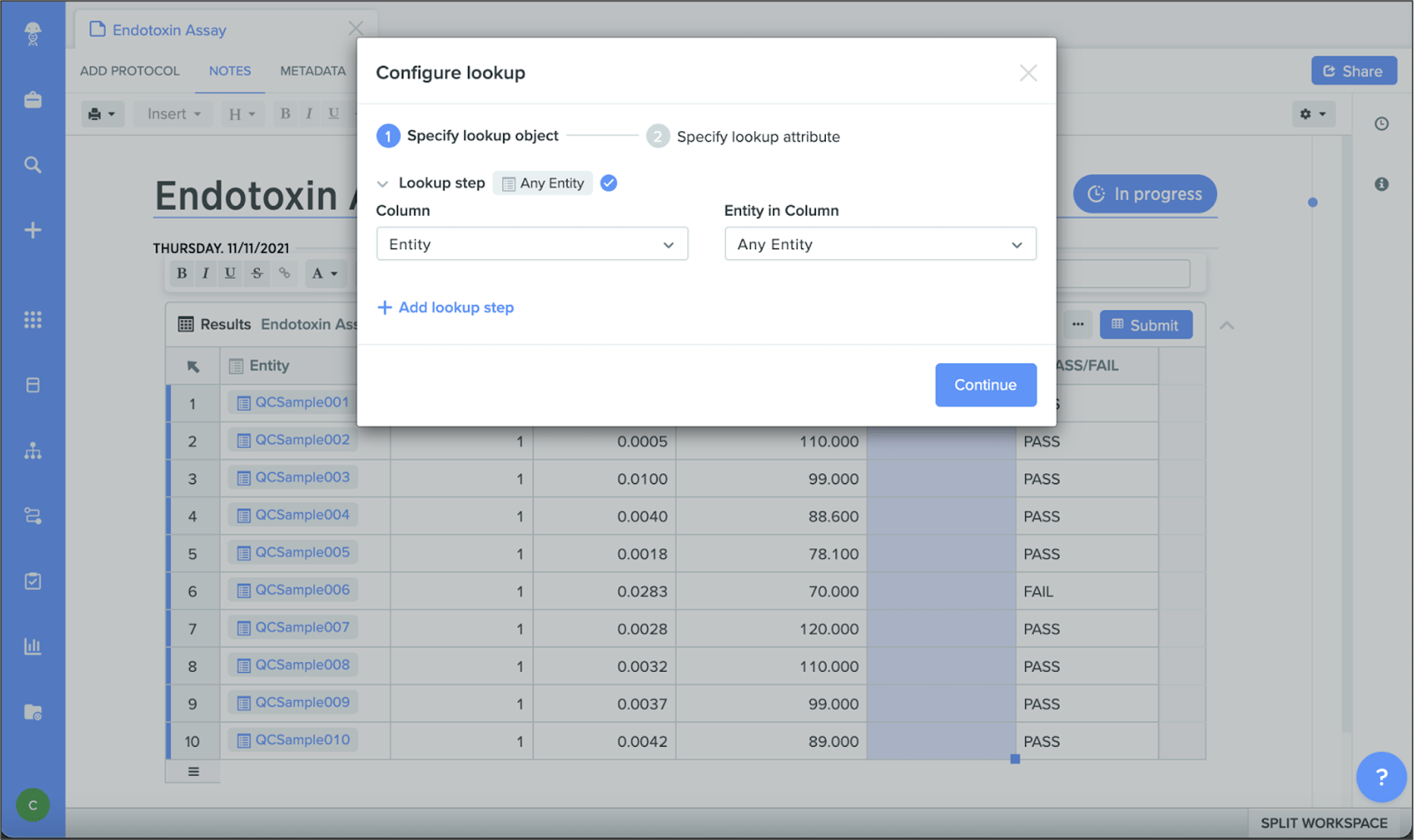
New results tables
New results tables in Notebook provide a more consistent user experience across tables in Benchling and streamline results capture for entities, plates, and containers. Combined with lookup tables and columns, results tables help you easily pull in information from worklists, spreadsheets, or other tables in Notebook. You can also better preserve the integrity of results with the ability to archive data and submit results tables.
Workflows
This summer, Benchling released a fully redesigned Workflows application to offer powerful task management, sample traceability, and operational intelligence. In this most recent release, teams can more easily link tasks related to a specific schema with task groups.
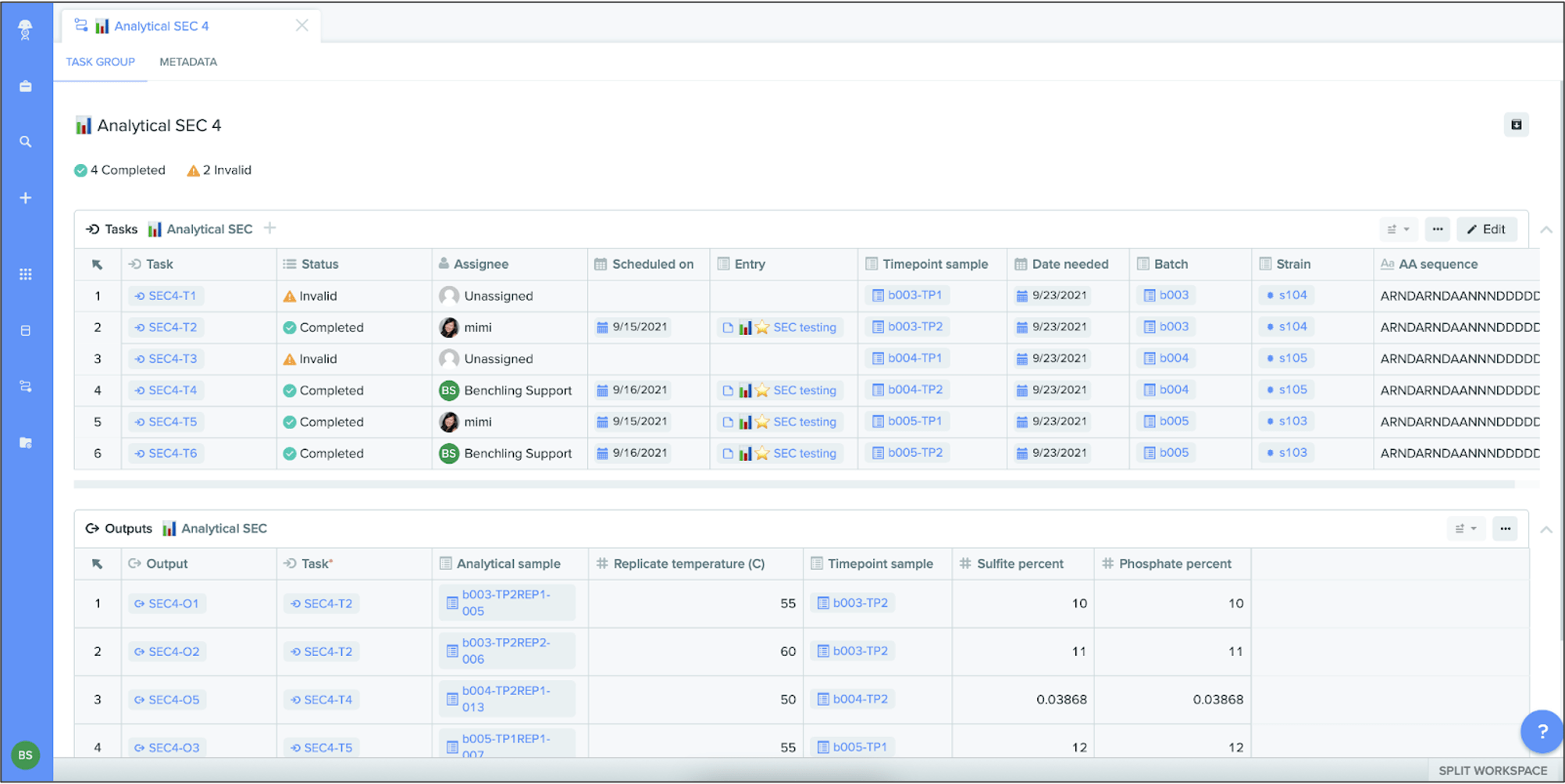
Task groups
In Workflows, users can create task schemas, which represent a specific job, such as “execute assay” or “request sample.” Now you can add one or more tasks of the same schema to a task group. These task groups help organize and track related jobs, while giving teams the flexibility to manage each task individually — including assigning, scheduling, completing, and tracking the status of each task. You can also add outputs for each task (via a Notebook entry or the API) that record the outcome of the task.
Insights
With Insights, you can visualize, query, and share Benchling data to help generate actionable intelligence about your R&D pipeline, experiments, and resource allocation. In this release, we’ve enhanced scatter plots with IC50 regression modeling, added heatmaps, and simplified how images display in Insights queries.
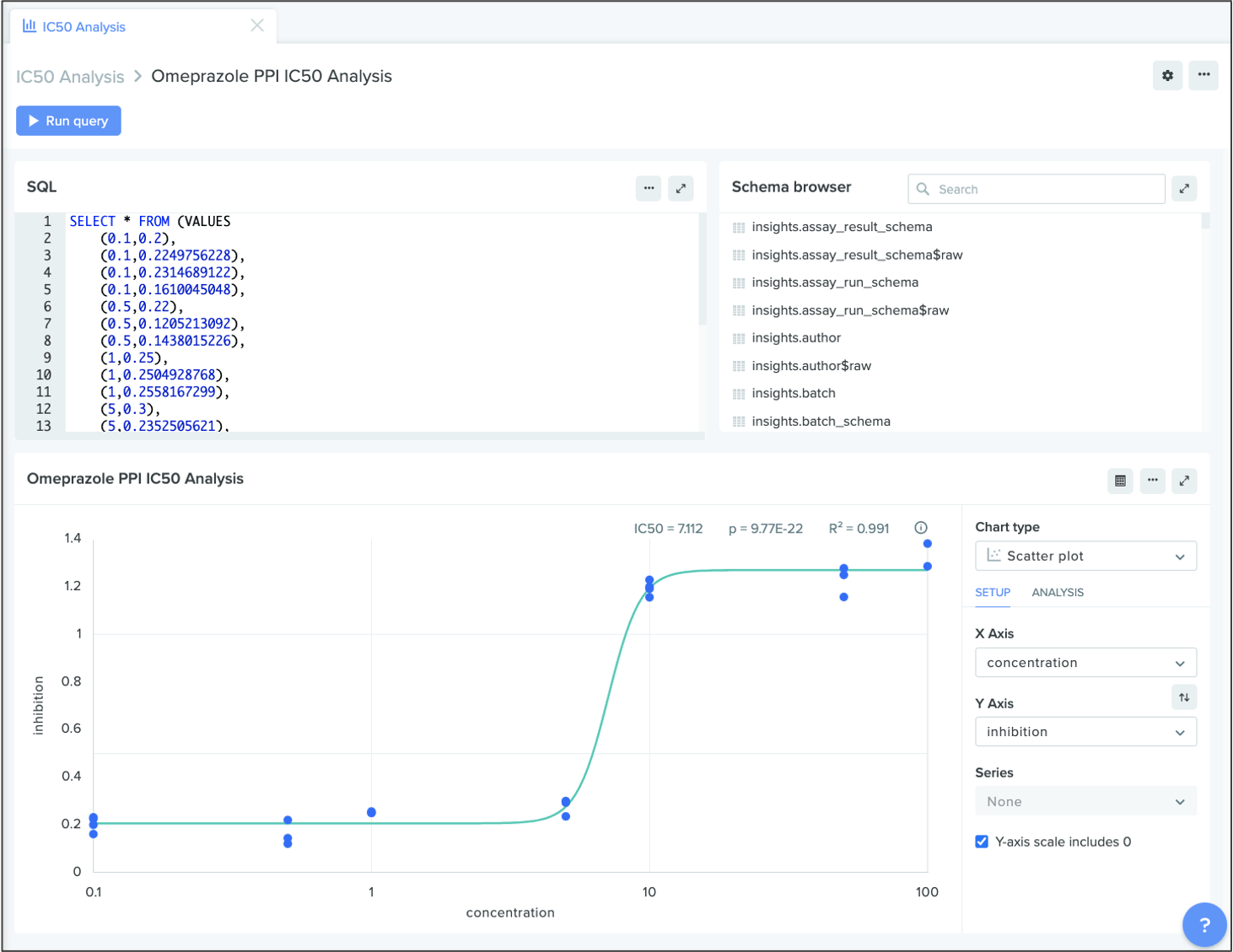
IC50 analyses on scatter plots
Last summer, we released support for linear regressions on scatter plots in Insights, making it easier to model multi-variable relationships in Benchling. In this release, we’ve extended scatter plot functionality to include IC50 chart types. These charts can illuminate powerful trends, such as the potency level of a certain drug during a toxicity assay. With just a few clicks, teams can generate these IC50 analyses and quickly visualize answers to key questions about R&D data.
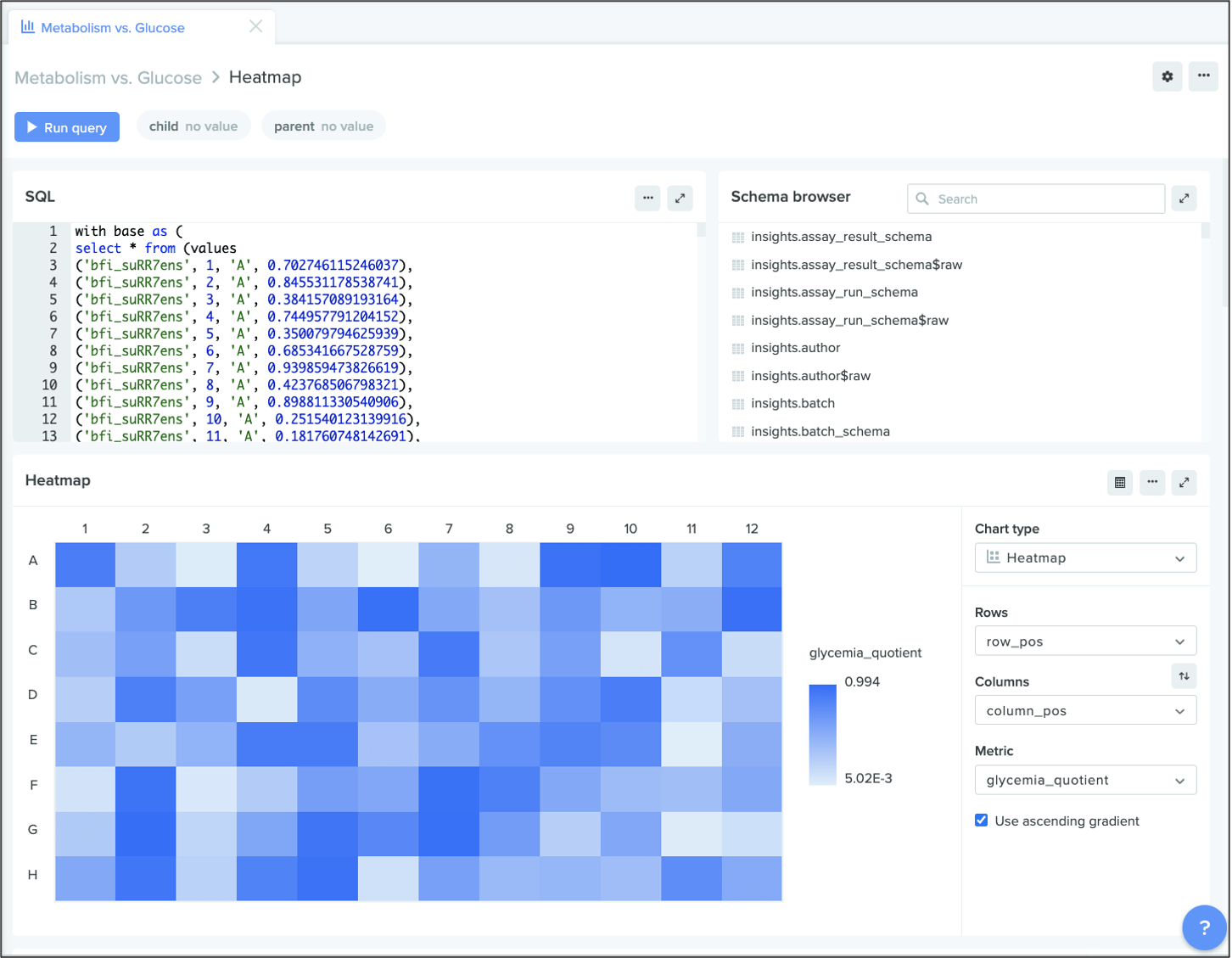
Heat maps
You can now create heat maps in Benchling Insights to better visualize your data. Heat maps are especially useful to compare and contrast different values within a plate. For example, if you want to understand the different fluorescent levels in a plate containing myosin protein samples, you can plot each cell’s fluorescence in a heat map to display results from dimmest to brightest.
Images displayed in Insights queries
In an Insights query, image blob links now display as images instead of image URLs.
Molecular Biology
Benchling Molecular Biology is an in silico design tool that supports a wide array of molecular modeling workflows, such as primers, digests, and CRISPR guides. In this release, we’ve improved key elements of the Molecular Biology user interface and made metadata from template alignments and external files more accessible.
Enhanced modified oligo functionality
We’ve made it easier to update and filter common modified oligo tasks.
Bulk update chemically modified oligos:whereas you could only use natural bases previously, now you can update modified oligos using CSVs with HELM or IDT syntax.
Monomer filtering:you can also refine and speed up modified oligo searches with new monomer filtering.
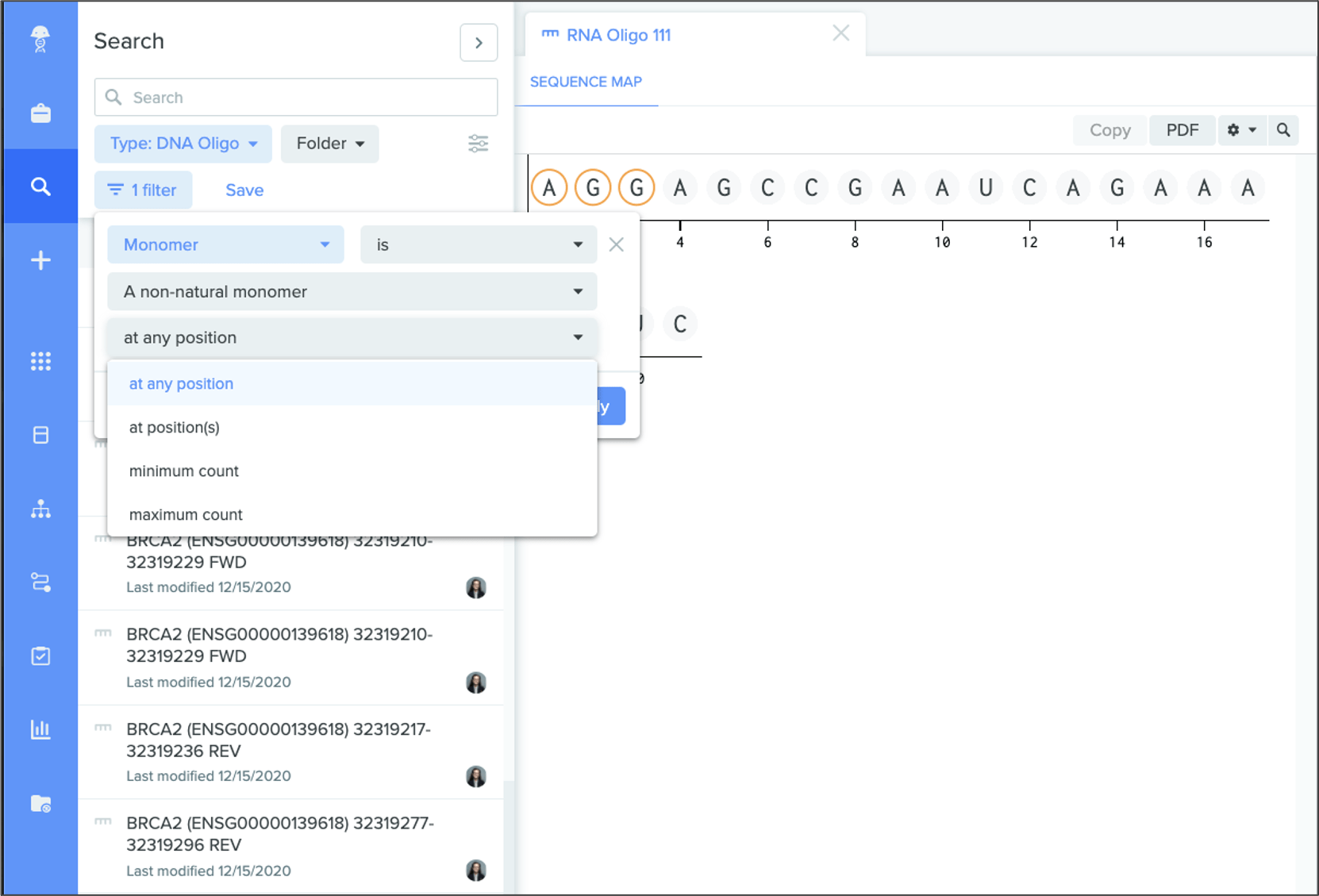
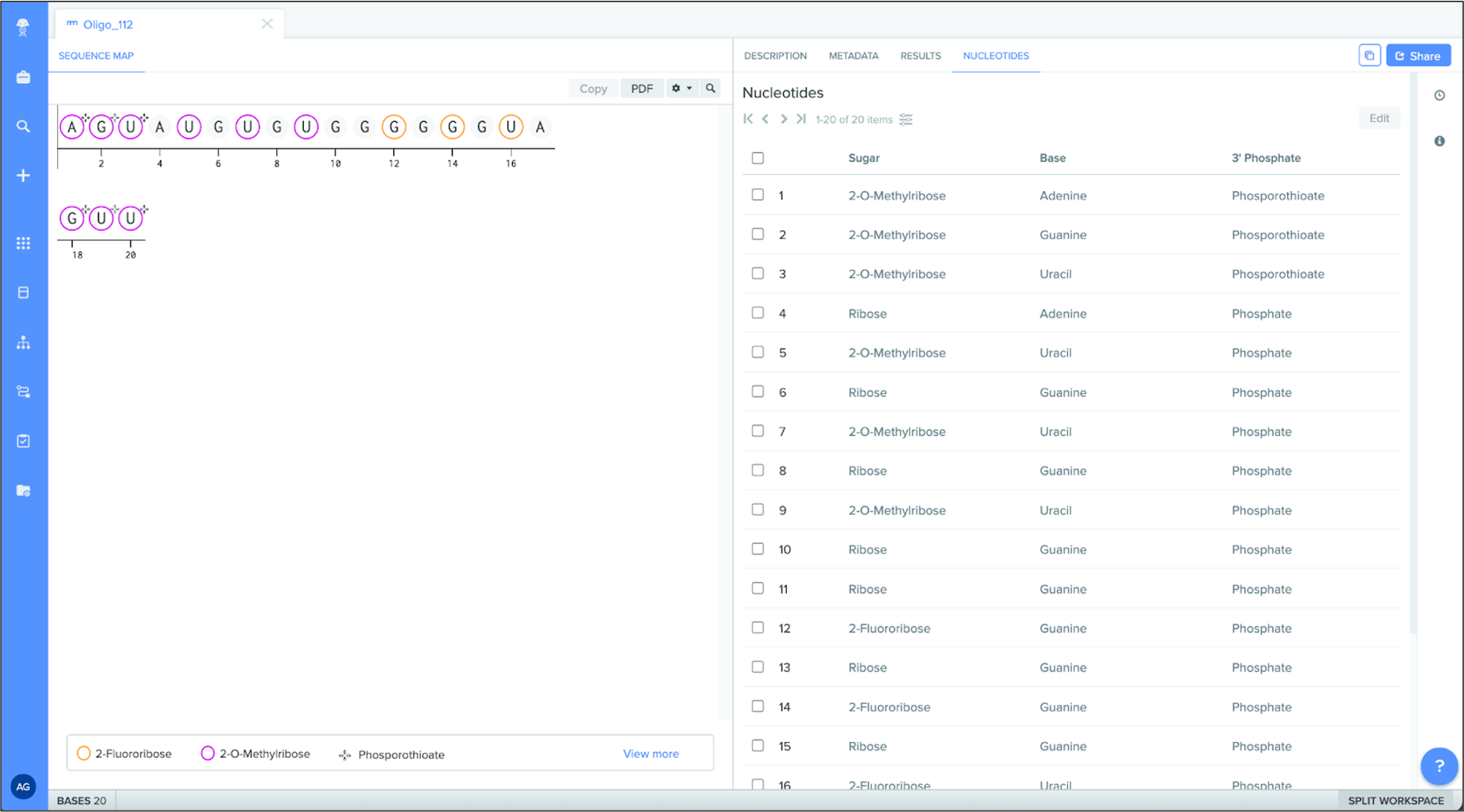
Customizable monomer colors
Registry admins can add custom colors to specific, non-natural monomers to better distinguish these molecules within an oligo or sequence.
Per-row statistics on template alignments
Template alignments now contain per-row statistics, such as length mismatches and pairwise identity, giving users quick access to key sequence metadata.
Improved visibility into external file metadata
When importing files from Geneious, GenBank, MacVector, and Snapgene, Benchling now populates the “annotations note” field with detailed file metadata. Previously, Benchling only brought over the file name and type.
Automatically create translation groups from Coding Domain Sequence (CDS) annotations
Upon import, Genbank files now automatically extract translation groups from CDS annotations. Users can immediately create an AA sequence from the entire translation group to produce a protein sequence that’s equivalent to the gene it encodes for.
Registry and Inventory
Benchling Registry and Inventory work together to power your experiments through connected data, whether that’s data on sequences, samples, or experimental results. With Benchling Registry, you can standardize, connect, model, and track samples across experiments. With Benchling Inventory, you can track vials, wells, batches and more, while linking each of these elements to relevant results. In this release, we’ve streamlined search, added new units of measurement for mixtures, and simplified key elements of the user interface.
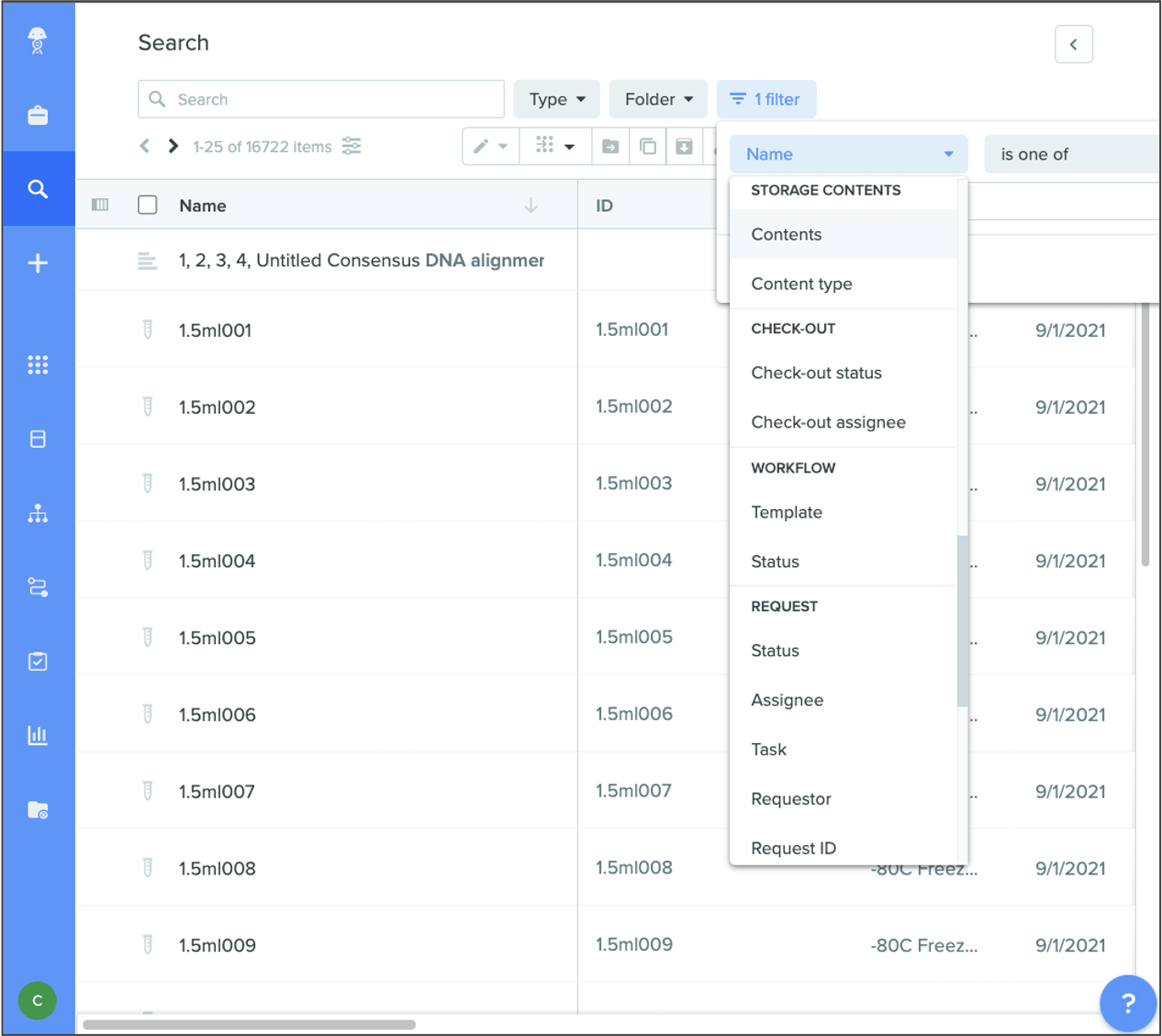
Search for container contents via global search
You can now search for specific items within a container by navigating to the “storage contents” filter type in global search, making information retrieval quicker. This replaces the “container contents” filter within Inventory search.
View full-sized images
You can now open a new tab to view full-sized versions of images that appear as preview images in Registry fields, schema fields, and Insights.
New units of measurement for mixtures
Mixtures now support the following units of measurement: moles per quantity (e.g., mol/L), micrograms per quantity, and kilograms per quantity. These new measurements allow teams to catalog more precise mixture data.
Enhancements to the user interface
We also enhanced how users create schemas and view information related to an entity.
Drop-down menu for schema selection:quickly navigate to and designate different schema types with the updated schema drop-down menu.
Improved schema naming:within the schema drop-down menu, the “Floating Point” schema type is now called “decimal” and the “Blob Link” schema type is now “attachment.”
Relevant items tab for entities:you can now view, search, and sort items related to an entity (like entries that reference the entity) in the “Relevant Items” tab in Registry.
Lab Automation
Lab Automation enables you to integrate lab instruments — like liquid handlers and plate readers — into Benchling to sync work across software and hardware, scaling experiments and increasing scientist productivity. In this release, we’ve enhanced compliance with audit logs and added new parameters for configuring runs.
Export Lab Automation audit logs
You can now export audit logs for runs, which contain information on the run’s description, action category and action, item and item type, related item, timestamp, and use. Additionally, the audit log field “Error files” has been renamed “Additional files.” Audit logs for run config changes are currently not available.
Automatic results archiving
Lab Automation now automatically hides archived results during runs.
Process samples and results simultaneously
You can now add results to the registration and transformation transfer bulk adapter to process these values in a single output step alongside samples.
Platform
The Benchling platform connects capabilities and data across all Benchling tools, providing a simple, unified user experience. In this release, we launched the Benchling Python SDK, added a dozen new APIs, and simplified key API permissions and policies.
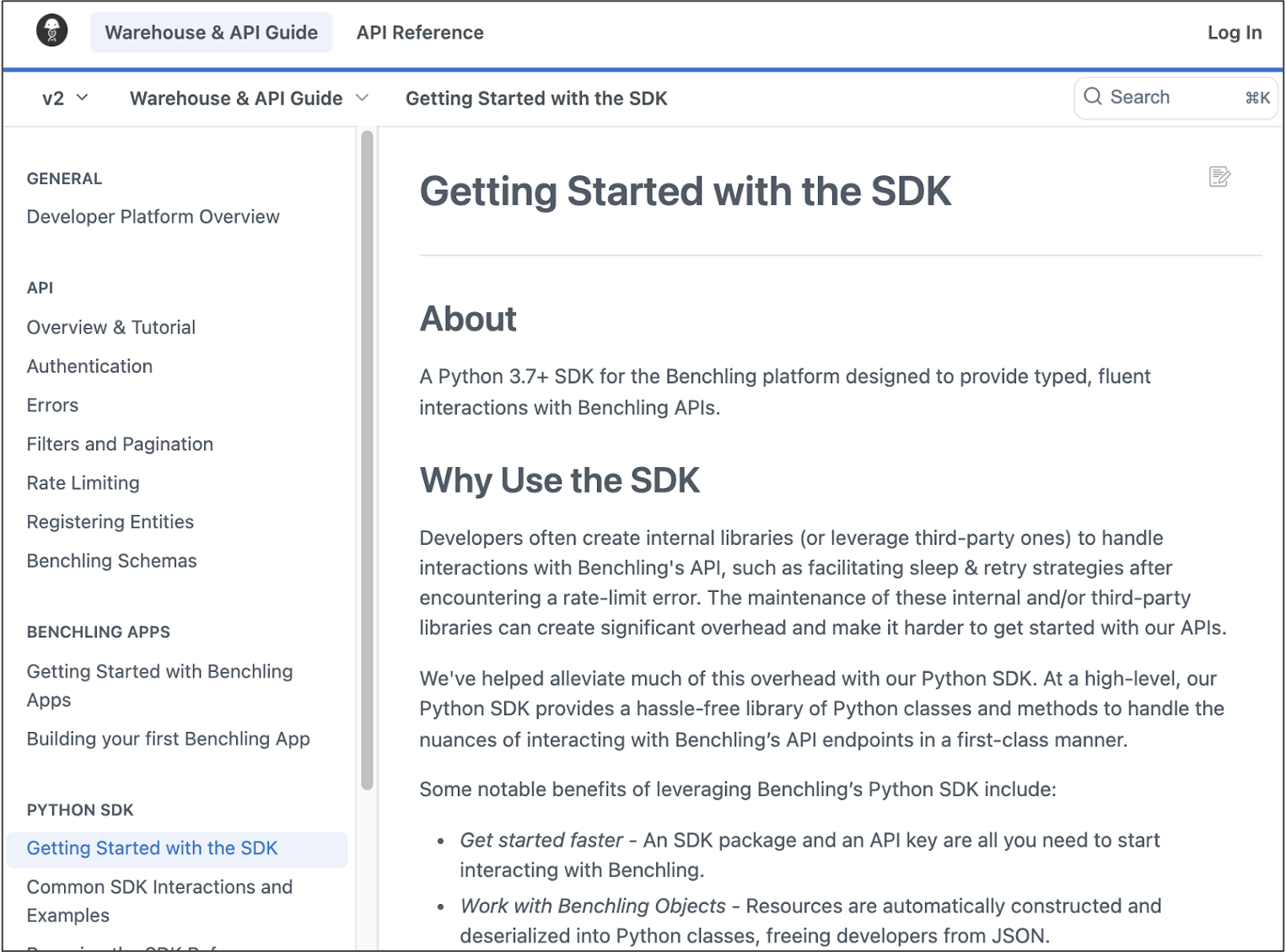
New Python SDK
Developers can now interact with Benchling APIs and develop on the Benchling platform using the Benchling Python SDK. This self-service resource makes integrations easy to set up and powerful to use, accelerating development timelines. With just the SDK package and API key, developers can begin to make API calls, integrate other systems, and easily maintain these integrations. Benchling takes care of connecting these systems, error handling, pagination, and re-entries off-the-shelf, which reduces the burden on IT teams.
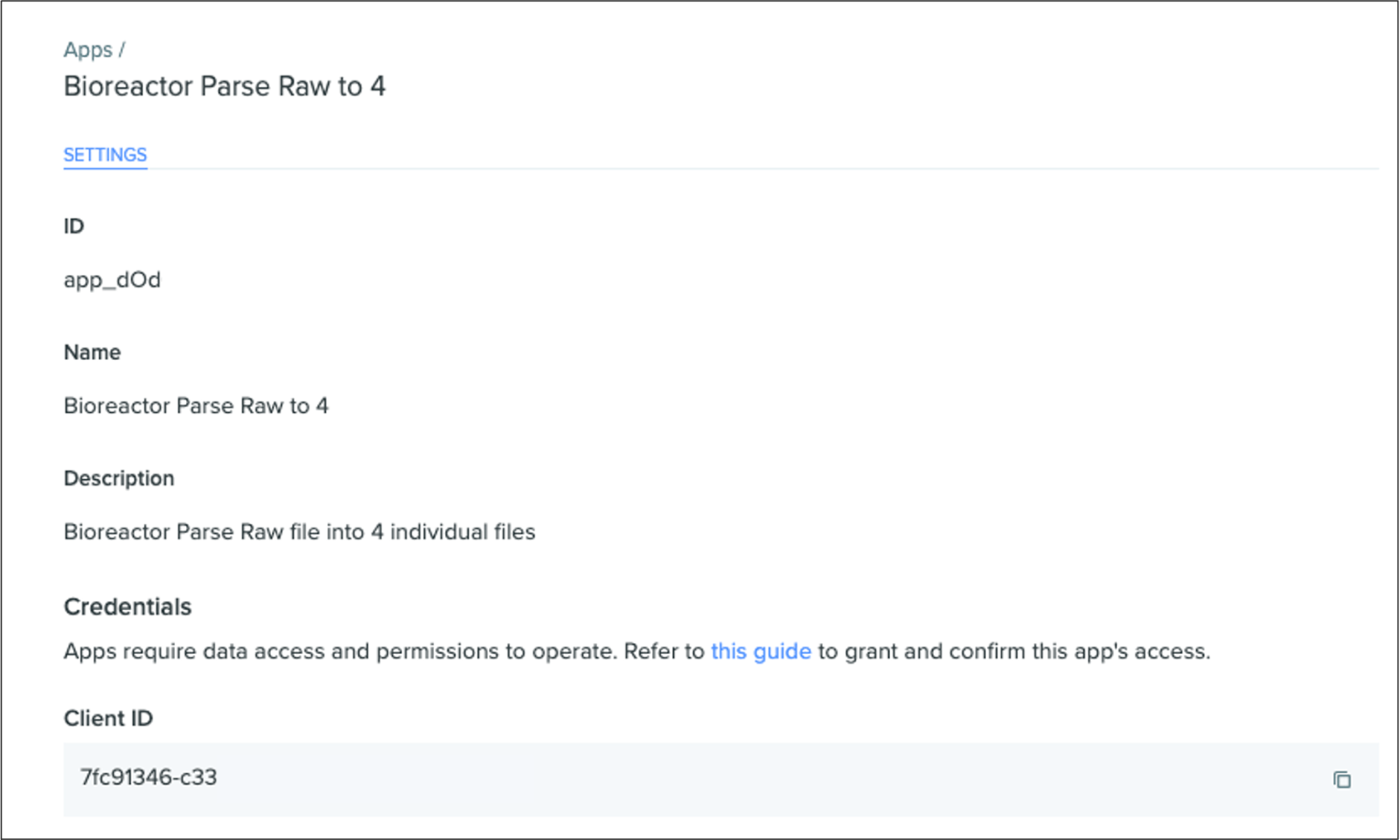
Benchling apps
Benchling apps provide a new integration framework within Benchling’s developer platform. With Benchling apps, developers and admins can develop, define, identify, manage, and permission application integrations independently from an individual developer’s credentials. Benchling apps improve how teams experience long-term data sharing between Benchling and other essential applications or instruments. These apps also provide standalone credentials and unique permissions for each integration. Keeping credentials independent from the developer helps organizations better manage app configuration and permissions, and gives teams greater visibility into app actions.
Improvements to the Benchling interface
Faster entry creation:add new entries from the Create menu in a single click. Previously, users had to navigate through multiple sections within the Create menu to add a blank entry.
New settings menu for Molecular Biology and other features:quickly navigate to Molecular Biology settings and feature settings directly from the “options” section of the settings menu.
Longer project lists:projects lists in Benchling now display 25 projects by default. You can also apply two new filters (project ownership and project status) to refine the list of projects.
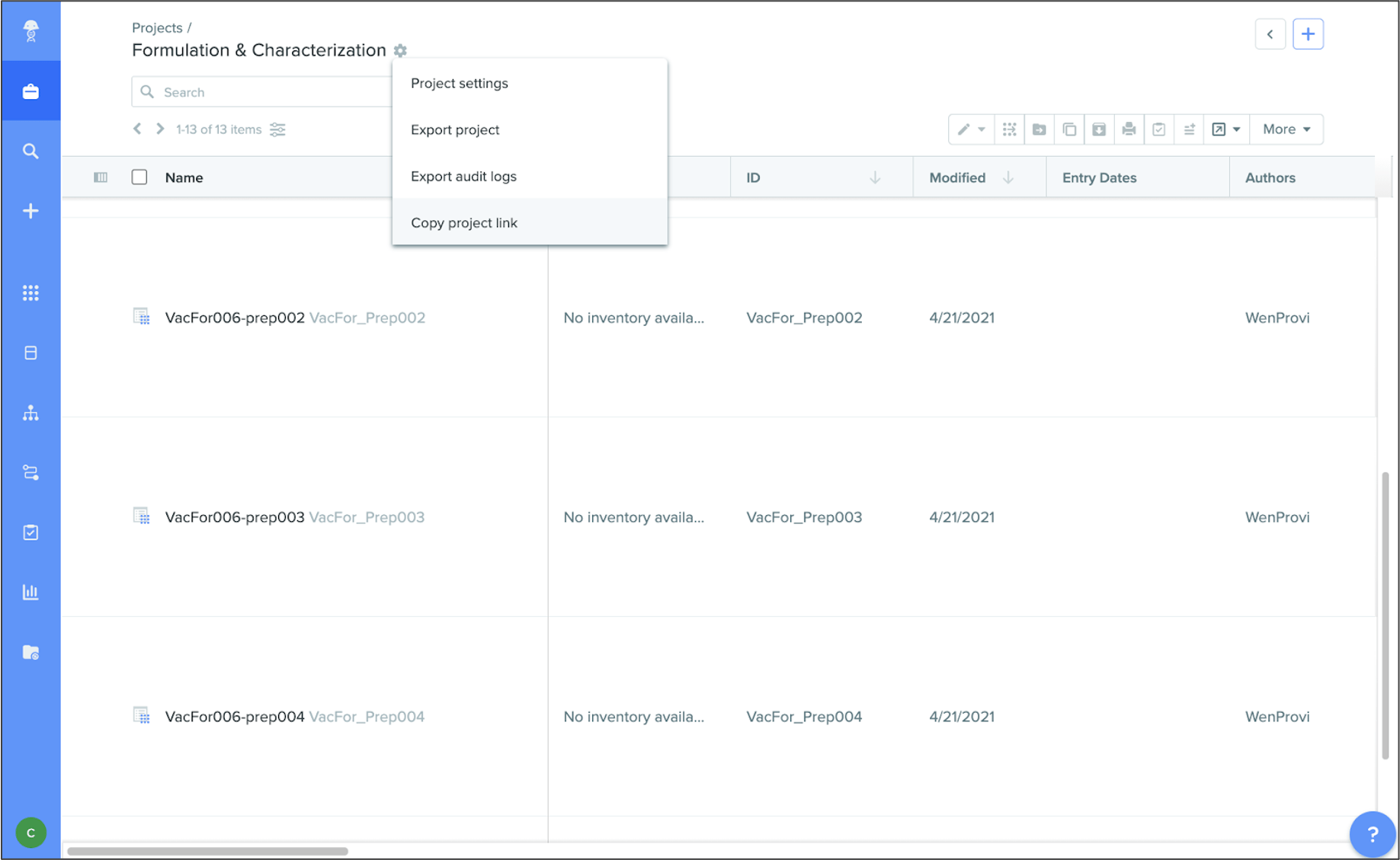
Copy links to projects and folders:copy links for projects and folders from the settings menu, and easily share these URLs with other users.
API updates
Benchling APIs are built to match the speed and agility of modern R&D, making it easy to extend Benchling into your broader digital ecosystem. We regularly enhance and add new APIs to further increase Benchling’s utility and expand the range of potential integrations.
This release, Benchling developers can leverage new APIs to:
List organism-specific codon usage tables
Create codon-optimized DNA sequences
Back translate AA sequences
Access mixture APIs via the public API
Expose the API ID of a reference sequence when executing an API call for DNA alignments
Filter on schemaId or displayId when using the entry list endpoint
Update custom entity authors when using the bulk updates endpoint
Clear input files on Lab Automation runs
Distinguish between entity types (AA sequence, mixture, oligo, etc.) when adding the new entityType field to entity responses
Filter users by an array of handles (part of the “Users” API)
Retrieve metadata for entry templates
Retrieve a specific box’s contents, including concentration and volume
Manage worklists through the API, which includes the ability to create, read, update, and delete worklists
Updated description for NamingStrategy API
Benchling API docs now include a more detailed description of the NamingStrategy API.
Better clarity on entity types during API calls
With the entityType field, developers can now distinguish between entity types (AA sequence, mixture, oligo, etc.) when these values are returned during an API call.
Improved control over personal API keys
The user settings page in Benchling now hides developers’ personal API keys so teams can screenshot and share these profiles without exposing sensitive information. Additionally, users can now copy API keys.
New policy statements for permissions API
You can now view policy statements for the “access policy” and “schema access policy” APIs, which detail permissions for a given policy.
Powering breakthroughs for over 1,300 biotechnology companies, from startups to Fortune 500s
