Discovering novel CRISPR effectors and a new RNA toolbox
Contributed by:
OMAR ABUDAYYEH AND JONATHAN GOOTENBERG
How do you expand the CRISPR toolbox?
CRISPR tools have mostly focused on the CRISPR-associated protein Cas9, a programmable DNA nuclease that serves as an important component of the bacterial adaptive immune response against foreign DNA threats. Cas9 has been repurposed for many types of applications, including targeted gene knockouts, transcriptional activation, epigenetic modifications, and much more. However, there has been much less emphasis on understanding what other programmable DNA proteins might exist, and whether there are even functionalities beyond just DNA targeting in the CRISPR universe.
In a previous study, we collaborated with Eugene Koonin's group, to understand the diversity of CRISPR systems [Shmakov et al 2015]. To uncover novel CRISPR-associated protein families, we decided to mine all sequenced bacterial genomes on NCBI, looking for unknown proteins that cluster near known CRISPR sites (Figure 1).
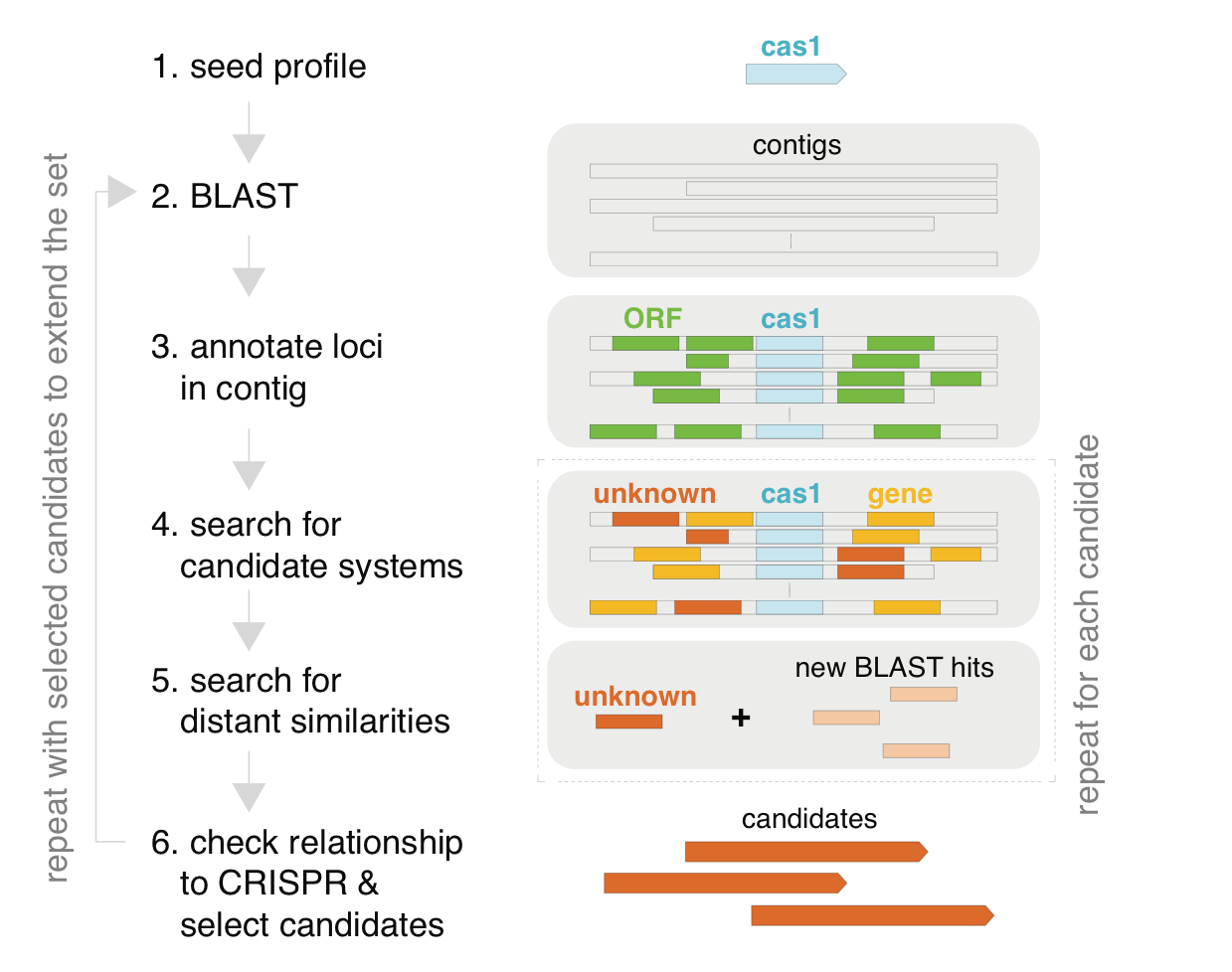
Figure 1. New Class 2 CRISPR systems can be discovered using a bioinformatics pipeline to detect effector genes near Cas1. For more, see [Shmakov et al 2015].
To our surprise, we found many families that no one had ever annotated or characterized, including a group of putative DNA nucleases similar to Cas9 called C2c1, and C2c3 (Figure 2), as well as Cpf1 [Zetsche et al], a previously detected but uncharacterized gene. Upon experimental characterization of these new enzymes, we were able to show that indeed they were programmable DNA nucleases with many unique features, highlighting the extensive diversity that exists in bacteria. Cpf1 in particular is quite different from Cas9 -- it has a T-rich PAM and requires only a crRNA, and Cpf1 has become an exciting new genome editing tool.
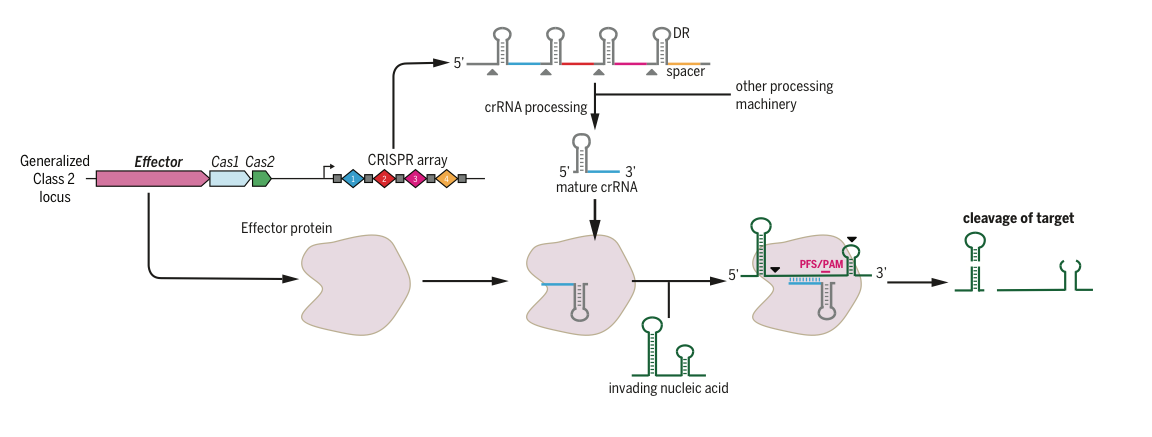
Figure 2. Class 2 CRISPR systems use a single programmable nuclease to target invading nucleic acids, such as phage genomes. Class 2 systems include Cas9, Cpf1, and C2c2.
These DNA targeting enzymes were not the end of our search, however. There was one curious family we called C2c2 that had no readily identifiable DNA nuclease domains. Instead, C2c2 had two domains associated with RNAse activity, called higher eukaryotes and prokaryotes nucleotide-binding (HEPN) domains (Figure 3). A single protein CRISPR system that only targets RNA has never been characterized before. We decided to try to understand exactly how C2c2 functions because of the unique biology and potential applications for RNA targeting.

Figure 3. The C2c2 locus contains a CRISPR array along with the Cas1 and Cas2 acquisition proteins. The C2c2 protein contains a pair of HEPN RNA-binding domains.
C2c2 is a reprogrammable RNA-targeting CRISPR protein
C2c2 is a unique CRISPR effector in two ways. First, unlike type III CRISPR systems that have a combination of DNA and RNA nuclease activity, C2c2 is the first CRISPR effector to naturally target RNA elements. Second, C2c2 is only a single protein, ~4,300 nt in length and 171 kDa in size. This is substantially smaller and simpler than the very large type III CRISPR effector complexes, which are comprised of many protein subunits (~13) and are ~427 kDa in size.
We started to study C2c2 by amplifying the LshC2c2 locus from the endogenous bacterial strain, Leptotrichia shahii, and cloning it into into pACYC184 (Figure 4).

Figure 4. The plasmid map for LshC2c2 locus in pACYC184.
We first showed that C2c2 is capable of defending against MS2 infection in Escherichia coli. MS2 is a ssRNA phage with no DNA intermediates in its life cycle, meaning that C2c2 must be interfering with the RNA in order to prevent infection. To verify and characterize the RNA cleavage activity of C2c2, we recombinantly purified C2c2 in E. coli. Upon incubation of the protein with a short RNA target and a crRNA complementary to a site in the middle of the target, we found that C2c2 cleaved the target at multiple sites in exposed ssRNA regions within the secondary structure. Moreover, it seemed that C2c2 had unique nucleotide preferences - the target site must not be flanked by a guanine (a feature termed the protospacer flanking site, or PFS) and that cleavage occurs at a nearby uracil (Figure 5). These results indicated that the enzyme could easily be reprogrammed to target any RNA by changing the sequence of the crRNA.
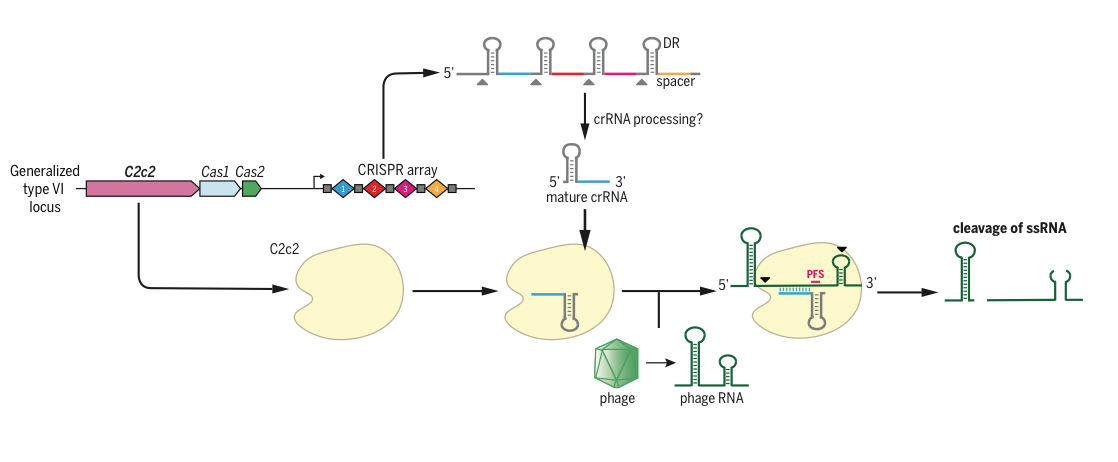
Figure 5. Type VI systems target RNA from invading phage, and cleavage is contingent on the presence of a Protospacer Flanking Site (PFS). Cleavage occurs in ssRNA regions.
Why do we need more tools to study RNA and how can C2c2 be useful?
Despite the wide variety of techniques for studying RNA biology, there are still significant gaps in our ability to either measure or manipulate RNA inside of cells. For example, it's possible to tag a messenger RNA with handles for recruiting fluorescent proteins to visualize how the RNA localizes and expresses in live cells [Bertrand et al 1998, Paige et al 2011]. However, this requires direct modification of the gene, and can't easily be scaled to multiple genes.
With a programmable RNA binding protein, you could direct GFP molecules to the site of specific transcripts in a cell and image them in real-time in a living cell. Additionally, while RNAi can be used to downregulate gene expression, it is neither specific or reliably efficient. C2c2 offers high specificity and has been shown to be very active for most crRNAs and targets tested, offering an alternative tool for gene knockdown. Beyond these applications, there exists a better need for controlling isoform expression, sensing specific isoform levels, regulating translation, and manipulating RNA trafficking.
To demonstrate the utility of C2c2 as a tool, we show specific targeting of RFP mRNA for degradation in E. coli by 20%-92% (Figure 6). We further show that the targeting of transcripts in E. coli to be specific with C2c2 activity being somewhat sensitive to single mismatches in the spacer region of the crRNA and highly sensitive to double mismatches.
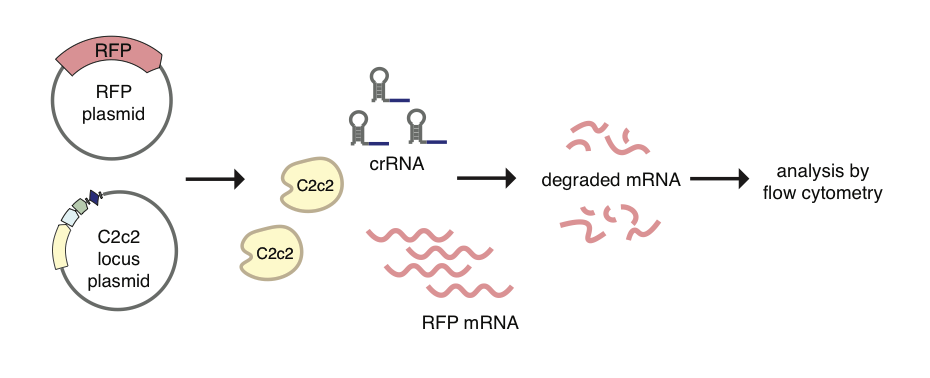
Figure 6. The C2c2 system can be re-engineered to target an mRNA inside We targeted Red Fluorescent Protein (RFP) and could observe a significant decrease in fluorescence when measured by flow cytometry.
Additionally, because many potential applications would only require binding, and not the cleavage activity, of C2c2, we generate catalytically dead versions of the enzyme and show specific binding of the enzyme to target transcripts. Overall, we believe C2c2 has many features to make it amenable for many exciting RNA applications in the future.
For a full description of how we found out C2c2, please see Abudayyeh, Gootenberg, Konermann, et al., Science, 2016