How to express CRISPR in your target cells
This tutorial is part 4 of 4 in the series “How to Create Knockouts Using CRISPR.” In the previous post, we talked about how to synthesize gRNA and Cas9 based on what cell line you will use. In part 4 of this tutorial, we discuss different delivery methods for your gRNA and Cas9. Use Benchling's free molecular biology tools to plan your own CRISPR experiment and design your own gRNAs here.
Introduction
By now you have designed your gRNA and synthesized the gRNA and Cas9 for creating gene knockouts. How should you deliver them to your cells? In the previous post, we provided a table to help you in selecting a delivery method based on the cell line you work with. Here we will discuss three standard delivery methods and analyze their pros and cons. For each option, we also provide a few further resources to conduct the experiment, including links to step-by-step protocols and plasmid maps.
Option 1: Lipofection
Lipofection is a lipid-based method to deliver DNA or RNA constructs to cells (Figure 1). The procedure is simple: You mix gRNA and Cas9 with lipid vesicles (liposomes) to form liposome-gRNA/Cas9 complexes. Then you add the complexes directly to your cell cultures. After 24 hours of incubation, the cells will start expressing gRNA and Cas9.
Here are the protocols to transfect gRNA and Cas9 to cells as plasmids or as RNAs.
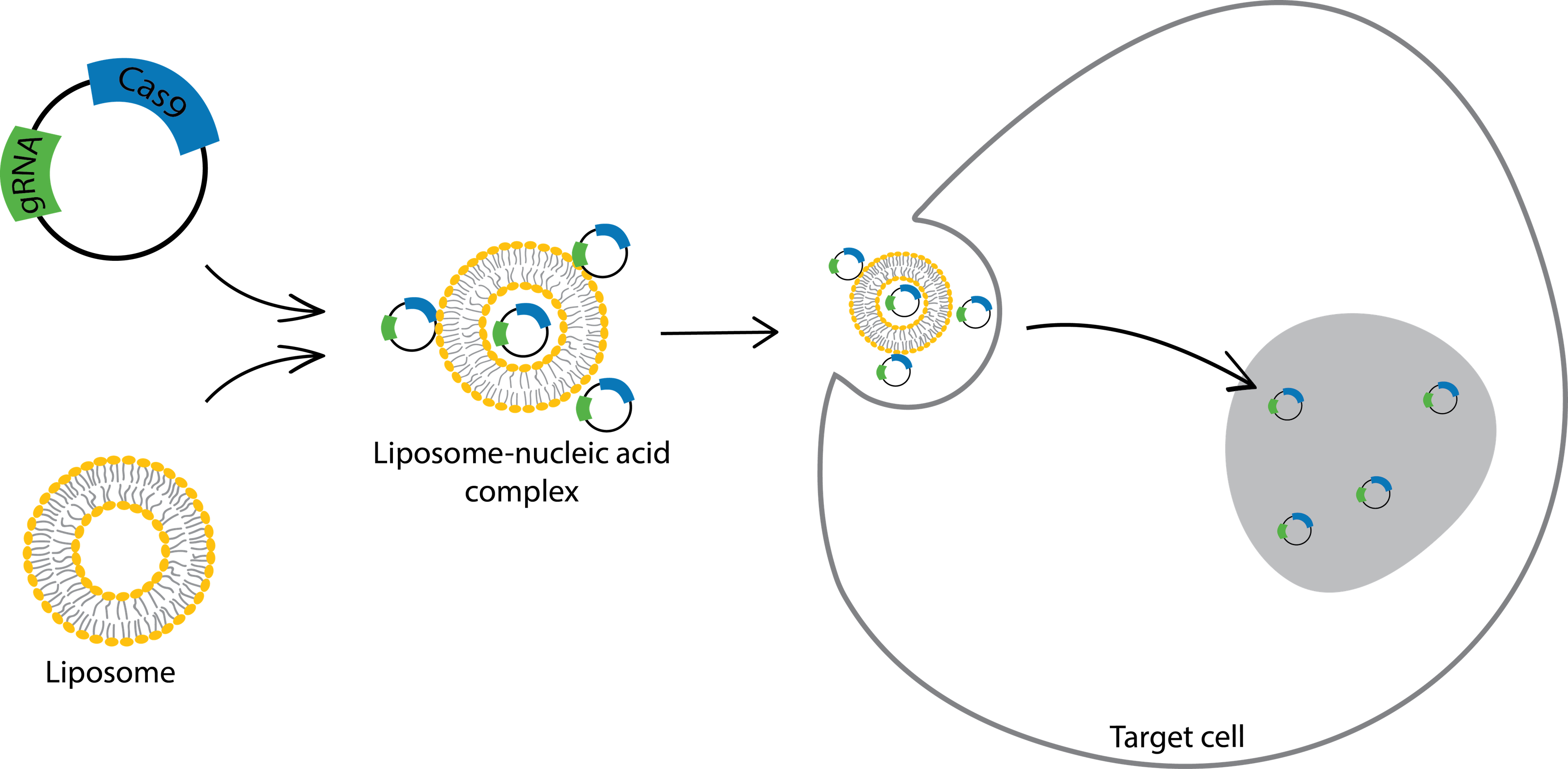
Figure 1: Deliver gRNA and Cas9 with lipofection. Liposomes are positively charged and can form complexes with negatively charged nucleic acids. These complexes are assumed to have net positive charges and allow cells to take up the plasmids easily.
Pros
The procedure is straightforward and does not require any special equipment.
Lipofection is flexible. You can use it to deliver gRNA and Cas9 in the form of either DNA or RNA. You can also adapt it for high-throughput systems.
Cons
Lipofection is not an effective delivery strategy for all cell lines. For example, if you use lipofection to deliver plasmids to human embryonic stem cells, you get almost 0% delivery efficiency. The table in this post provides the lipofection delivery efficiency for some common cell lines.
Tips
To maximize the delivery efficiency, use cells with low passage numbers and that are at 70%~90% confluence.
To optimize the delivery efficiency, make sure the quality of your nucleic acids is high. To estimate the quality of nucleic acids, you can measure the ratio of their absorbance at 260 nm over their absorbance at 280 nm (A260/A280). Good-quality DNA usually has an A260/A280ratio of around 1.8; good-quality RNA has an A260/A280ratio of around 2.03.
Having trouble delivering gRNA and Cas9 to an easy-to-transfect cell line? Try altering the ratio of nucleic acid to liposome, the density of cells, and the volume of transfection reagent.
Do not add antibiotics during lipofection because it will cause cell death.
Option 2: Electroporation
Electroporation is a procedure that uses an electrical pulse (300~400 mV for less than a millisecond) to create temporary pores on the cell membrane that DNA or RNA can enter (Figure 2).
Here is a protocol for electroporation written by Alex Ling, a graduate student in Dr. Alberto Martin’s lab.
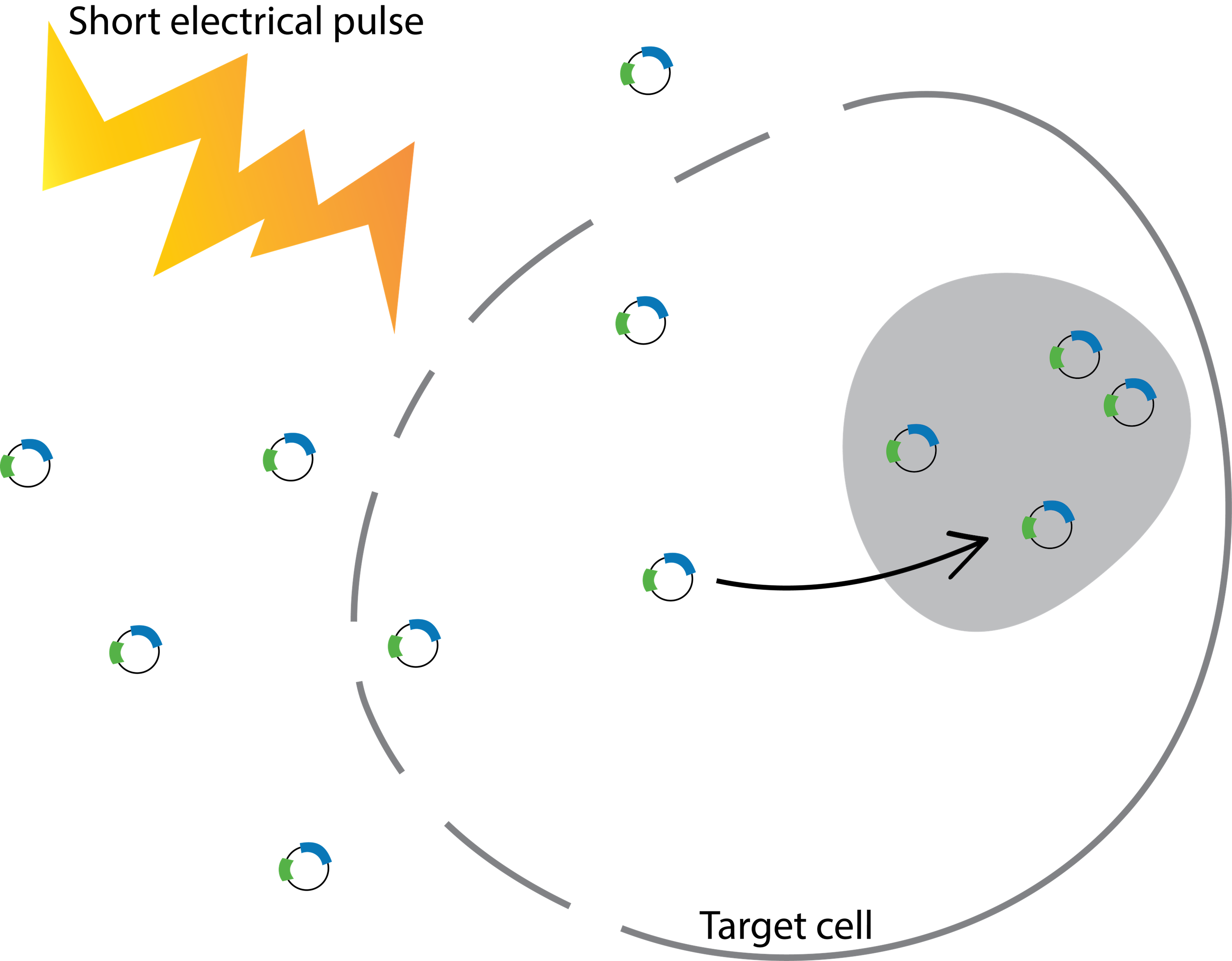
Figure 2: Deliver gRNA and Cas9 with electroporation. Electroporation exposes cells to a short electric pulse, which disturbs the cell membrane and forms temporary pores. The pores are conductive, so charged molecules, such as gRNA and Cas9, can cross the cell membrane and enter the cell.
Pros
It takes only a few minutes to use electroporation to deliver your DNA or RNA to cells. Therefore, once you optimize the conditions for electroporation, you can transfect gRNA and Cas9 to a large number of cells in a short period of time.
Cons
You need to either build or purchase an electroporation apparatus for electroporation. If you only use it occasionally, it might be hard to justify the effort.
Since electroporation involves applying high electric pulses to cells, it can lead to substantial cell death. Therefore, it is important to test and optimize the duration and strength of the electric pulse for each type of cell to minimize cell damage or death during the procedure.
Tips
Salts and proteins can lower electroporation efficiency, so it is important to purify and dissolve your nucleic acids in water or TE buffer.
The cells are fragile after electroporation, so it is essential to immediately add a recovery medium to the cells to keep them healthy.
Option 3: Lentiviral transduction
You can also deliver your gRNA and Cas9 with lentiviral vectors, which are derived from a subclass of Retroviruses called human immunodeficiency virus 1 (HIV-1). You first transfect plasmids in 293T cells to assemble functional lentiviral particles. You then can harvest the lentiviral particles and transduce them to your target cells. After the lentiviral particles enter a cell, the viral RNA, including your gRNA and Cas9, is reverse-transcribed, imported into the nucleus and inserted into the cell’s genome at a random location (Figure 3). You can find the protocols for lentiviral transduction from the Broad Institute or the Trono lab.
Here are the plasmid maps for pCMV-dR8.74psPAX2 (a packaging plasmid) and pMD2.G (a pseudotyping plasmid).
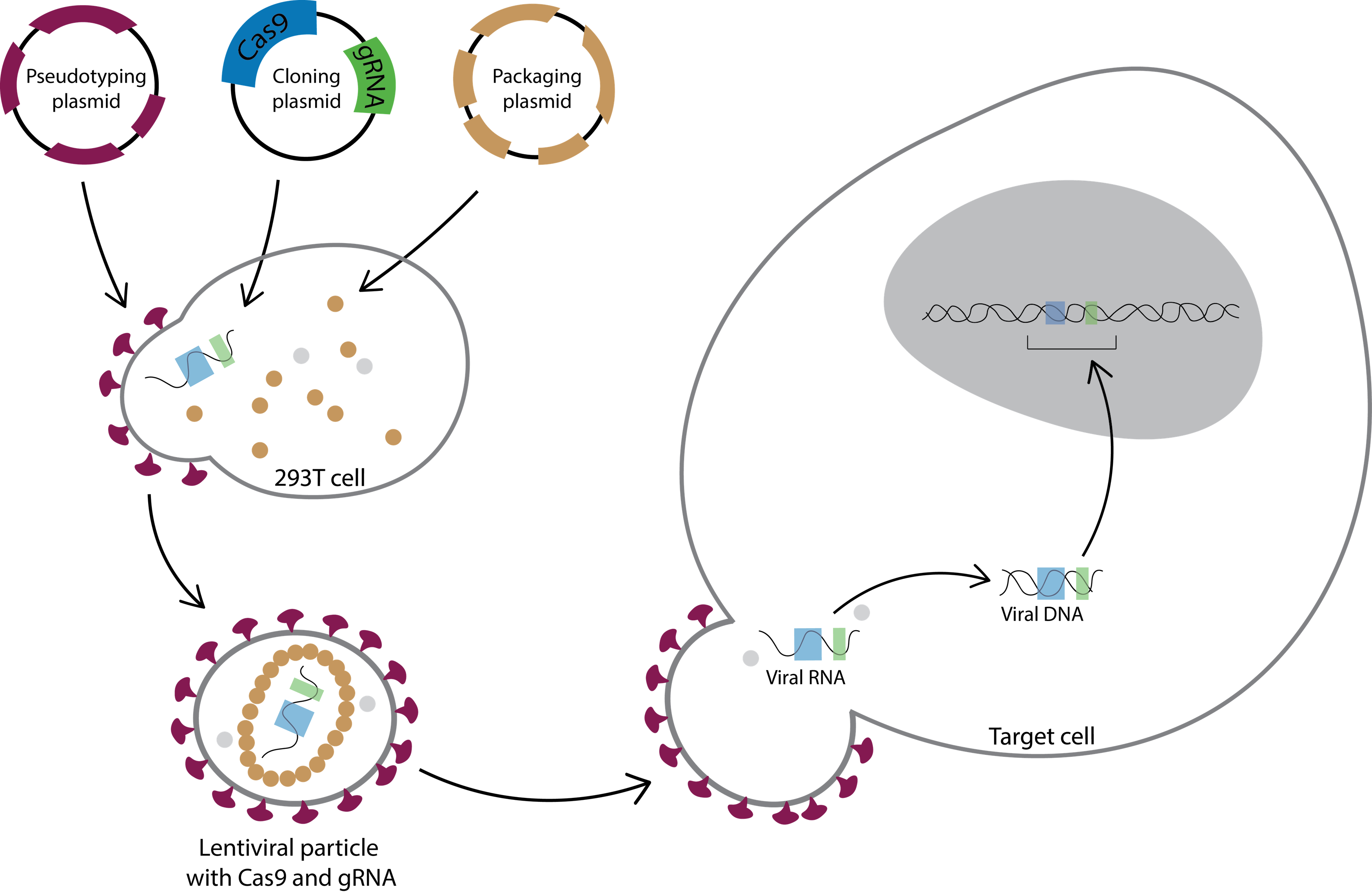
Figure 3: Deliver gRNA and Cas9 with lentiviral transduction. Lentiviral particles are typically produced by transfecting three three plasmids to 293T cells: (1) a pseudotyping plasmid; (2) a cloning plasmid; (3) a packaging plasmid. The pseudotyping plasmid encodes an envelope protein (purple); the lentiviral cloning plasmid includes psi packaging sequence, your gRNA and Cas9; and the packaging plasmid contains capsid proteins (brown) and reverse transcriptase (gray). The 293T cells express the plasmids and produce lentiviral particles. After you harvest the lentiviral particles, they can then be transduced to target cells. Once they are in the cells, the reverse transcriptase (gray dots) reverse transcribes the viral RNA genome to a viral DNA genome. The viral DNA genome then is imported into the nucleus and incorporated into the cell’s genome at a random location. Your cells will stably express gRNA and Cas9 24-48 hours after lentiviral transduction.
Pros
The delivery efficiency is high even in the most hard-to-transfect cell lines, including differentiated neurons.
Cons
The procedure is laborious and takes around two weeks to produce and transduce the lentiviral particles.
Depending on where the viral genome is integrated into the host genome (which is impossible to control), it might affect important functions of the cell. If you are concerned about the problem, you might want to try adeno-associated virus (AAV). You can learn more about the mechanism of AAV transduction here.
Tips
To achieve high viral production, you should prepare 293T cells within 24 hours of transfection.
After you generate the lentiviral particles, aliquot the viral stocks, so you don’t have to freeze-thaw the stock often.
If the cells look unhealthy after lentiviral transduction, try to lower the amount of lentiviral particles you use or change the growth medium 4 hours after transduction.
What’s next?
From the previous post, we covered which method will give you the best delivery efficiency in some of the most common cell lines. In this post, we have explained how to implement the delivery method. In the next post, we will discuss functional assays you can use to detect insertions and deletions (indels) in your cells.
Want to use Benchling's CRISPR tools at your company?
References
Felgner, Philip L et al. "Lipofection: a highly efficient, lipid-mediated DNA-transfection procedure." Proceedings of the National Academy of Sciences 84.21 (1987): 7413-7417.
Felgner, Jiin H et al. "Enhanced gene delivery and mechanism studies with a novel series of cationic lipid formulations." Journal of Biological Chemistry 269.4 (1994): 2550-2561.
Sambrook, Joseph, Edward F Fritsch, and Tom Maniatis. Molecular cloning. New York: Cold spring harbor laboratory press, 1989.
Neumann, Eberhard et al. "Gene transfer into mouse lyoma cells by electroporation in high electric fields." The EMBO journal 1.7 (1982): 841.
Potter, Huntington, and Richard Heller. "Transfection by electroporation." Current Protocols in Neuroscience (2011): A. 1E. 1-A. 1E. 11.
Naldini, Luigi et al. "In vivo gene delivery and stable transduction of nondividing cells by a lentiviral vector." Science 272.5259 (1996): 263-267.
Powering breakthroughs for over 1,300 biotechnology companies, from startups to Fortune 500s
