How to Upgrade from Vector NTI to Benchling
Modern molecular biology deserves modern software
For information on importing Vector NTI files into Benchling, follow the instructions at the bottom of this post.
In the past decade, molecular biology has evolved tremendously, but sequence design software has stagnated. Most sequence design tools are point solutions that weren’t built for modern scientific workflows or cross-team collaboration. Such tools ultimately can’t meet the full needs of complex R&D workflows, since they don’t meaningfully integrate with broader informatics infrastructures. As a result, R&D organizations struggle to tie downstream assay data back to sequences and can’t fully leverage the data they’re producing. This regularly slows down research and and introduces risks into process development and scale up.
Within today’s life science R&D organizations, molecular biology is high-throughput, collaborative, and organizationally complex. To meet these needs, companies need much more than a desktop-based point solution. They need cloud-based software unified with a broader informatics platform, so they can transfer context across teams and trace from sequences down to the results of individuals candidates.
In our whitepaper "How to Upgrade from Vector NTI to Benchling" we discuss how you can digitally transform your molecular biology with Benchling's modern, cloud-based platform.
How to import your Vector NTI files into Benchling
Before you import
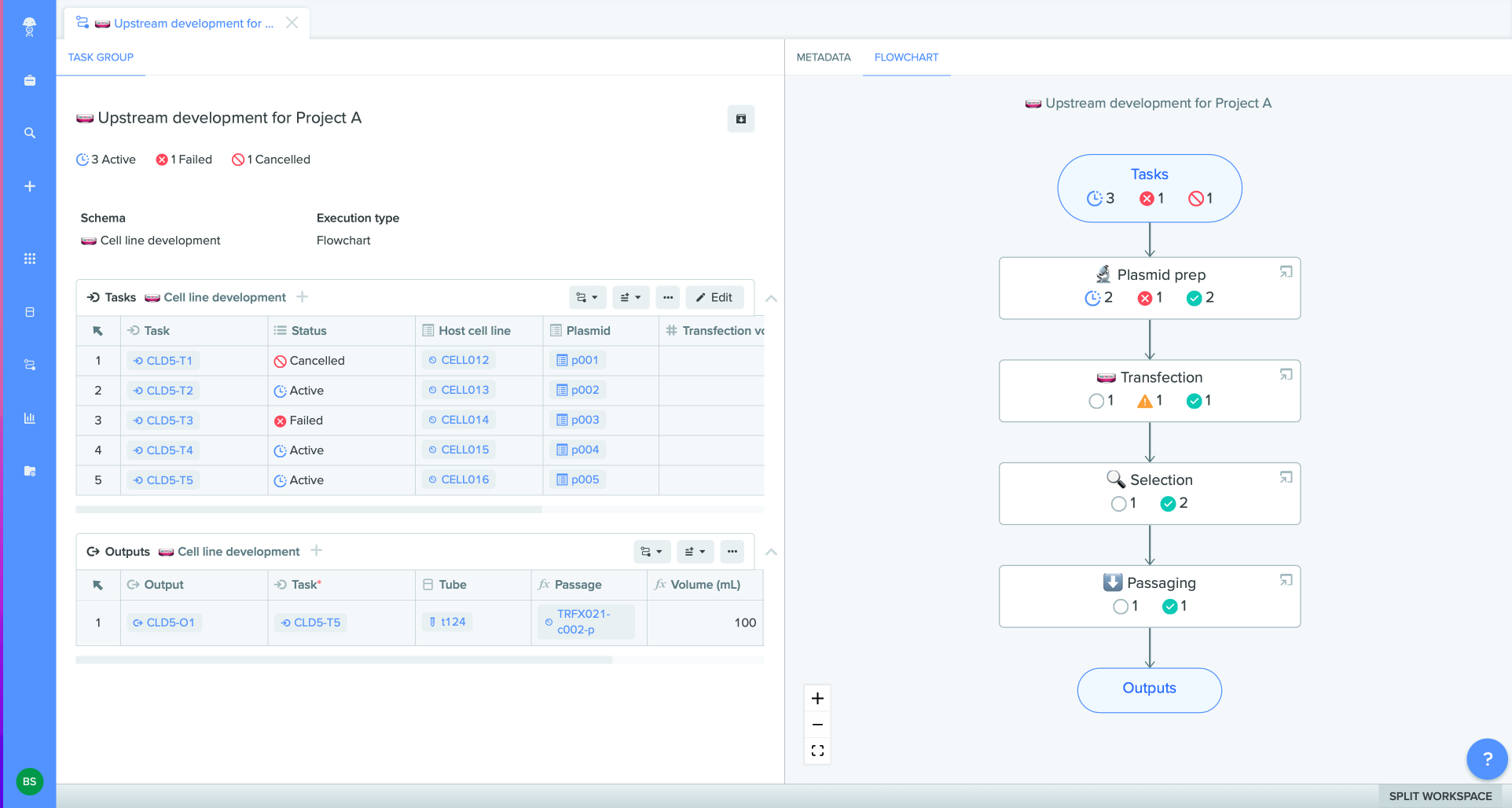
Benchling is capable of hosting complex project structures, similar to Vector NTI. Before you begin importing your Vector NTI files, make sure to establish your ideal project and folder structure in Benchling. You can do this by clicking the projects tab, then the + button at the top of the left-hand panel to create your project. Click the+ button once more and select the option to create a new folder. Once you have your project, folder, and any subfolders created, you are ready to import your files into Benchling.
Preparing your Vector NTI files for import
Benchling can support a host of file formats, including .ma4 and .pa4. When you import your files into Benchling, data such as annotations, trace qualities, and primers that are associated with a particular sequence will be carried over. Rather than downloading all of your files individually, download them as an archive file. Once your files are downloaded to your computer, they are ready to be imported into Benchling.
Drag and drop your files
Open the destination folder for your file. Click the + button, then select whether you are importing DNA, amino acids, or oligos. In the pop-up window, make sure you are in the Convert Files tab, and simply drag and drop your archived file from your computer into Benchling.
NOTE:The original file will still exist on your computer after this action. Dragging and dropping your file into Benchling makes a copy of the original file to be stored in Benchling.
After your files are imported
Benchling will then show you all of the sequences within the archived file that you have imported. Click on an individual sequence to see the associated annotations and descriptions of the file type and sequence.
You can highlight multiple sequences to take a bulk action, such as move them to a new folder, open them in Benchling, or set a tag by which to search for these files in the future. If you need to move a file or multiple files to a different folder later on, click on the → button near the top left to access Benchling’s expanded window where you can select all of the files that belong in a different folder and move them in bulk.
Have questions?
If you have any questions about importing files into Benchling, you can reach out to our team via the in-app help window. To access this feature, click on the ? button in the lower right corner of your Benchling window.
Powering breakthroughs for over 1,300 biotechnology companies, from startups to Fortune 500s