Introducing Benchling Model Hub: The infrastructure layer for scientific AI
Today we're launching Benchling's Model Hub. Scientists can now run the best predictive models without standing up the infrastructure themselves. The predictions they generate are automatically fed back into the underlying R&D record, informing the next experiment and closing the loop between the wet and dry lab.
Model Hub is a dedicated space inside Benchling where scientists can discover, run, and track scientific AI models, connected to their experimental data and R&D workflows. It's accessible via a new icon in the Benchling navigation bar, giving it a permanent home alongside the rest of your scientific work.
For anyone interested in adding computational discovery to their work, this means no more infrastructure overhead, no context switching between tools, and no manual effort to connect model outputs to experimental records. You select your inputs, choose your models, run predictions or generative analyses, and get back structured results with a full audit trail, all inside Benchling.
The last mile problem in scientific AI
Scientific AI models have made remarkable progress, but deploying them inside a real R&D workflow still requires significant time, infrastructure, and technical overhead.
For most organizations, running models requires compute provisioning, DevOps work, API integrations, and ongoing maintenance. Teams that go this route can spend months grappling with GPU availability, open source repos, and more before a scientist ever runs a prediction.
Model access is central to our AI Scientist vision. We started by embedding structure prediction tools directly into our Molecular Biology suite, giving scientists a way to run models like AlphaFold, Boltz-2, and Chai-1 alongside their sequences. Customers told us they wanted more ways to bring in data, more visibility into what models were available, and the ability to run predictions across massive batches of sequences at once, not one at a time.
Models and partnerships
We’re launching Model Hub with an initial curated set of trusted models. We’ll continuously expand the range of available models, bringing in both open source advances and proprietary models from partners as the scientific AI ecosystem moves forward.
New open source models
OpenFold 2 and OpenFold 3-preview are open source reproductions of AlphaFold2 and AlphaFold 3, respectively, developed by the OpenFold Consortium and the AlQuraishi Lab at Columbia University. Benchling also recently joined the OpenFold Consortium to further our commitment to supporting the community at large.
Protenix is ByteDance Research's structure prediction model, capable of modeling proteins and small molecule ligands as part of a single complex. It brings an additional option for teams working on multi-component biological systems where understanding the full interaction picture matters.
Proprietary model access
Continuing in our pursuit of democratized model access, we will be adding access to proprietary models in the coming weeks, including a partnership with Boltz PBC, the team behind Boltz-2 and BoltzGen. Benchling also intends to make Lilly TuneLab available on the platform, as previously announced.
New features
Model Hub now has a dedicated home in Benchling, accessible via a new icon in the left navigation bar. From there, scientists can browse the full model library, kick off prediction runs, and track results in one place.
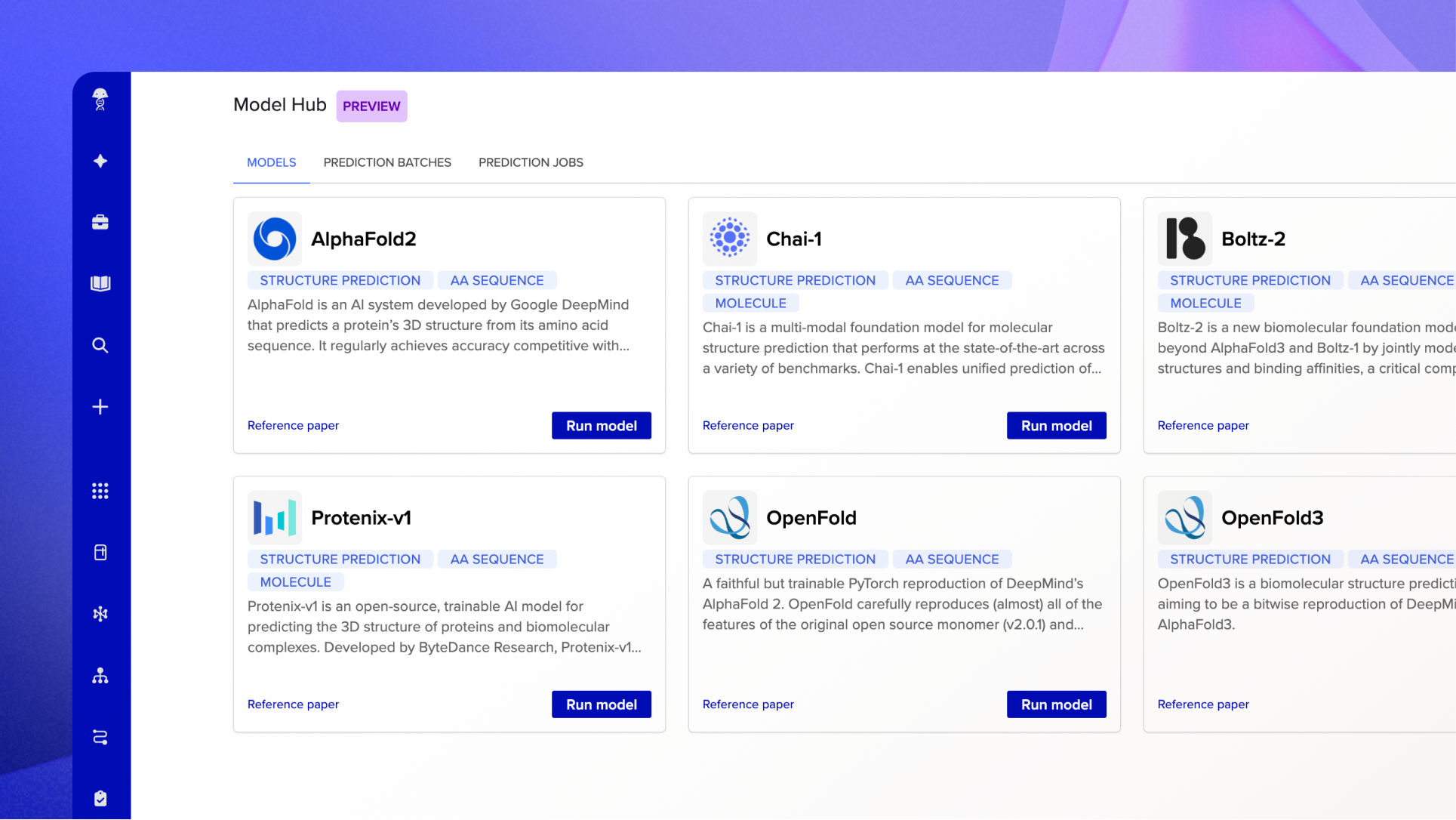
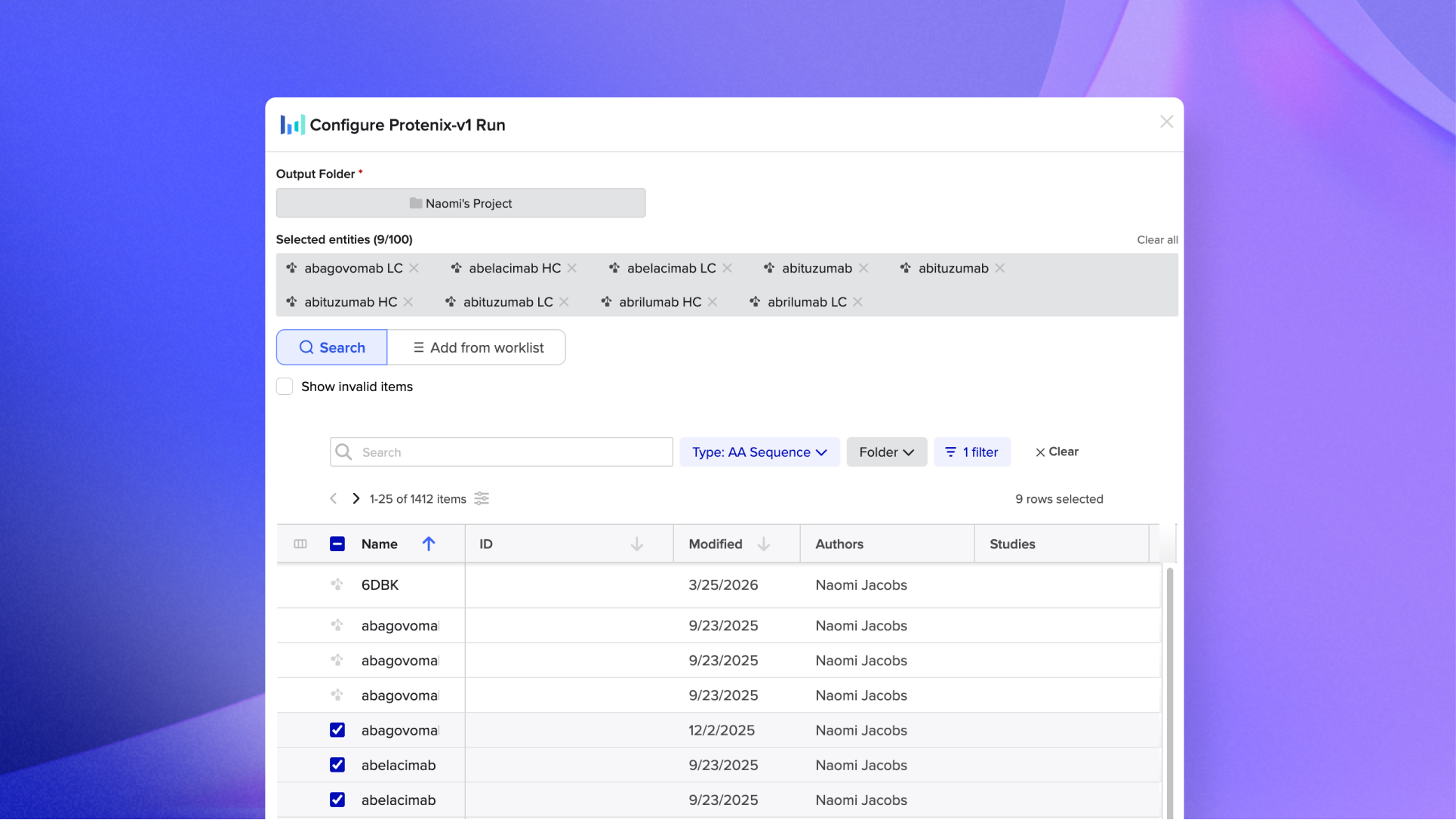
Batch Predictions allow you to select a large set of sequences and submit them all in a single run. Select structure predictions across a hundred proteins, choose your model, and run. Results come back as an organized prediction batch, ready to review and act on.
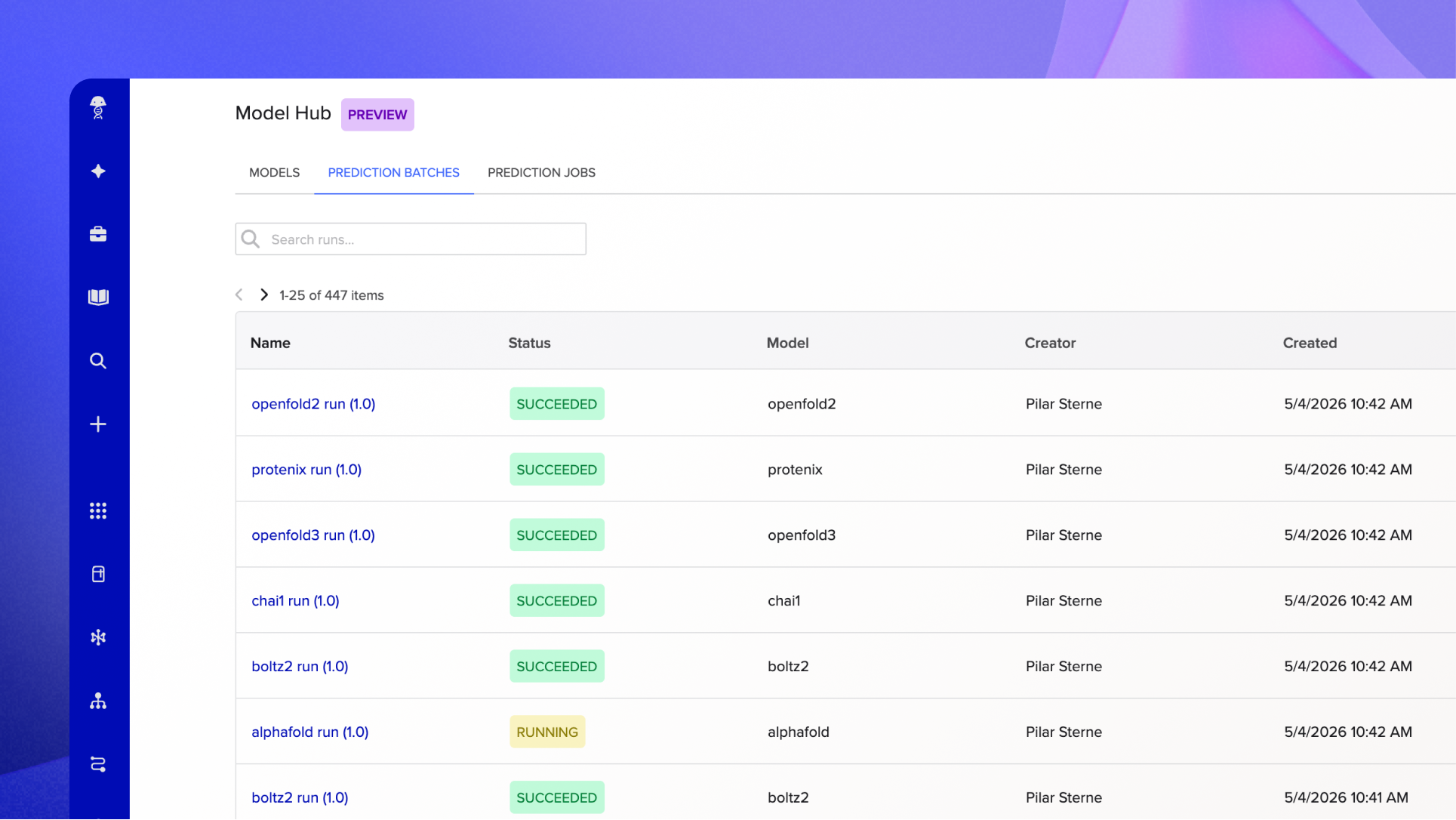
Prediction tracking gives you a centralized view of everything you've run: every model, every input set, every result. It's the record of your in silico work, organized and accessible, rather than scattered in local folders. You can review outputs, trace results back to the sequences or experiments they came from, and share findings with collaborators.
Faster execution, more predictions per credit: Benchling has overhauled the underlying infrastructure behind our Scientific Models product. Newer GPUs provide a significant boost in speed. With this update, you can now run 4X the structure predictions with the same credit allocation.
MSA (Multiple Sequence Alignment) support improves prediction quality for structure models that benefit from evolutionary context by incorporating alignment data alongside your input sequences. We run GPU-accelerated MSAs, cutting what has traditionally been one of the slowest steps in the prediction pipeline down to a fraction of the time.
Try it today
Already a Benchling customer? Model Hub is available in your account now. Look for the new Model Hub icon in the navbar to get started (see here to learn more).
New to Benchling? Sign up at Benchling.ai to explore Model Hub with a sample dataset and see what's possible.



