Sickly Sweet – How Utilisation of Sugars Affects the Disease Progression of Streptococcus pneumoniae

This article was contributed by:
Vikrant Minhas
Graduate Student, Paton LabThe University of Adelaide
Disease Burden and Strain Complexity
S. pneumoniae is a gram-positive bacterial pathogen that infects multiple sites of the body, causing diseases such as pneumonia, otitis media, bacteraemia and meningitis. These diseases result in the death of an estimated 1 million children and in billions of dollars in health care costs annually.1 Despite this massive mortality and morbidity, S. pneumoniae is commonly carried asymptomatically in the nasopharynx, with a small subset of pneumococcal strains capable of spreading deeper into the body and causing disease.1 Of these, individual strains can differ greatly in their capacity to cause disease. For example, some are only capable of infecting the middle ear, while others can only invade into the lungs. Major factors that allow S. pneumoniae to cause disease are known; in particular, we know that the capsule that surrounds the bacteria helps to evade the host immune system.2 Nonetheless, the mechanisms that dictate where pneumococcal strains spread remain poorly understood.
The poor understanding of pneumococcal disease can be largely attributed to the species' vast genetic diversity: at least 98 capsular serotypes and more than 12,000 clonal lineages or sequence types (ST). This complexity has made it difficult to determine the mechanisms that allow certain strains to infect specific sites of the body.3 With ever increasing antibiotic resistance and only partial coverage of current vaccines, we need to understand the mechanisms of pneumococcal disease progression to help design optimal vaccines and therapeutics.4,5
Clonal Comparisons to Determine Virulence Mechanisms
This is where the Paton Lab in the University of Adelaide's Research Centre for Infectious Diseases comes into the picture. We are attempting to understand disease progression of S. pneumoniae by focusing on closely related strains within the vast sea of pneumococcal genetic diversity. Previous studies from our lab have shown that clinical clonal isolates of S. pneumoniae, belonging to the same serotype and sequence type (ST), spread to different sites in mice; these sites corresponded to their original isolation sites in humans. For example, serotype 3 ST180 and serotype 14 ST15 isolates from the blood of human patients invaded the lungs and/or the blood of intranasal challenged mice. Clones isolated from the ears of patients could not invade the lungs, but they instead spread to the ear and/or brain.6,7
Why can strains from the same clonal lineage stably infect different sites of the body? To investigate this, we performed whole genome sequencing on the blood and ear isolate pairs from the serotype 14 ST15 and serotype 3 ST180 clonal lines.8 Although members of the same serotype and ST type are genetically very similar, they have acquired distinct genetic changes such as single nucleotide polymorphisms (SNPs) or insertions or deletions. We aligned the genomic sequences of each blood and ear isolate pair with the goal of identifying any changes that could be responsible for their varied disease phenotypes.
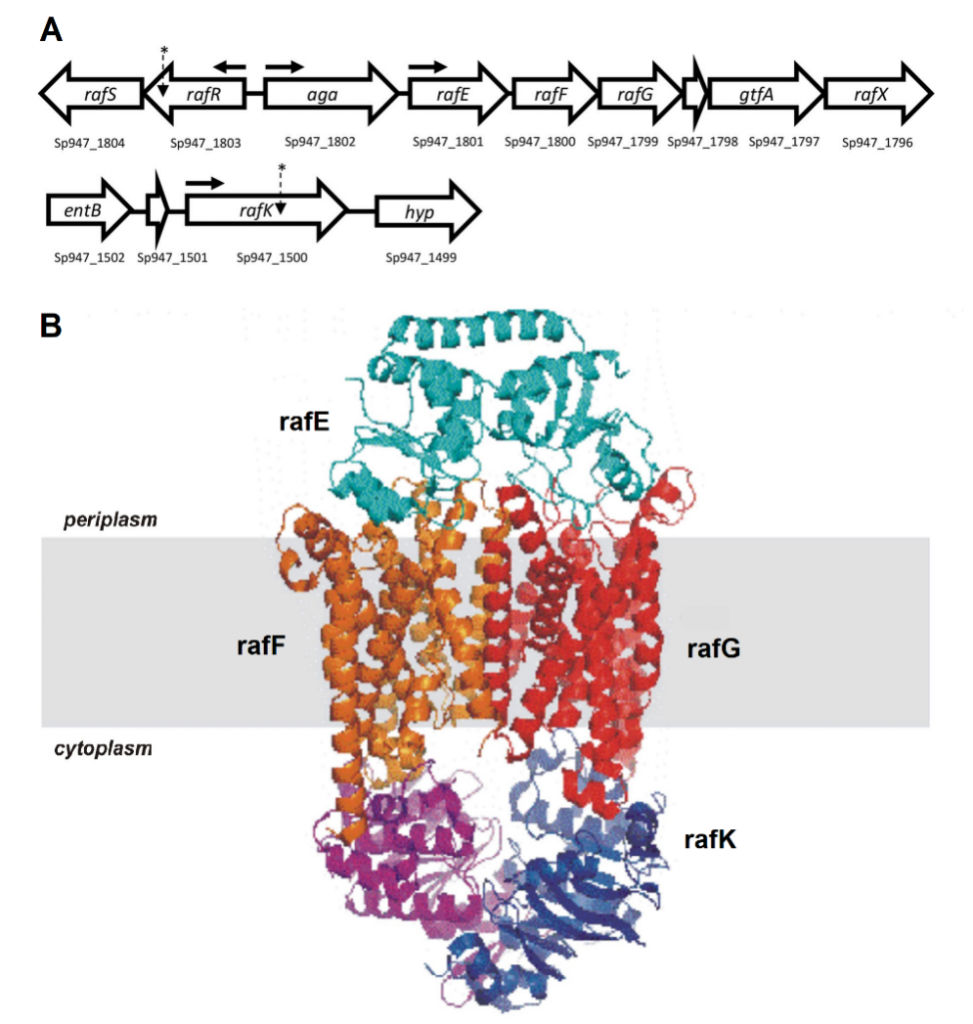
Figure 1. Raffinose uptake operon. (A) Genetic loci encoding raffinose uptake and utilisation in S. pneumoniae. The numbers below each gene refer to the locus tags in the serotype 14 ST15 947 genome. The locations of SNPs in serotype 14 ST15 and serotype 3 ST180 isolates are indicated with asterisks; horizontal arrows show the locations of promoters. (B) Visual representation of the proteins that make up the ABC transporter of raffinose.
Raffinose Utilisation Dictates Pneumococcal Disease Progression
Genomic comparisons identified SNPs in the gene rafR for the serotype 14 ST15 clonal pair, and in the gene rafK for the serotype 3 ST180 clonal pair. Both of these genes are part of the raffinose operon, which encodes an ATP binding cassette (ABC) transporter and multiple enzymes responsible for the uptake and metabolism of the trisaccharide sugar raffinose (Figure 1). RafRis the regulator of this operon while rafKacts as the ATPase required by the ABC transporter for raffinose uptake.
S. pneumoniae is a strictly fermentative bacteria; it relies solely on carbohydrate metabolism for energy, growth and subsequently virulence, as sugars provide the simplest and most readily available forms of carbon and energy.9 Hence, we decided to investigate the effects of the SNPs in the raffinose genes.
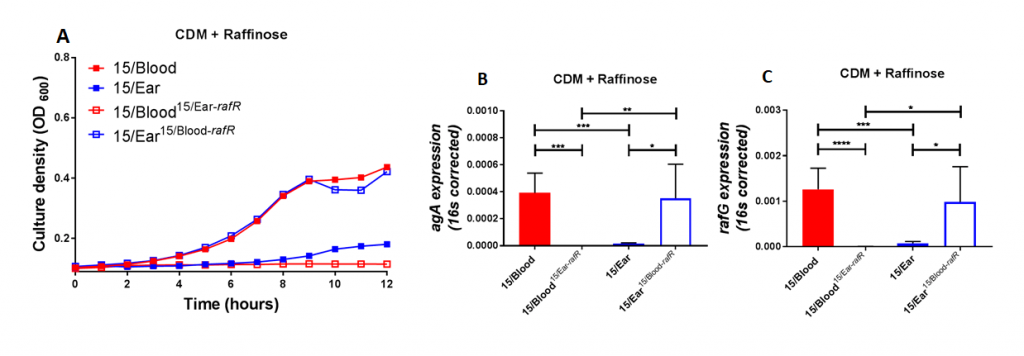
Figure 2. Growth phenotype and raffinose pathway gene expression in serotype 14 ST15 blood and ear isolates. (A) Growth of blood isolate, ear isolate and respective rafR swapped mutant in CDM+Raf was monitored by OD600 for 12 h. Data are mean OD600 ± SD from triplicate assays. (B&C) The indicated strains were grown in CDM+Glc to an OD600 of 0.2, washed and resuspended in CDM+Raf, and then incubated at 37°C for a further 30 min. RNA was then extracted, and levels of aga (B) and rafG (C), mRNA were analyzed by qRT-PCR using 16S rRNA as an internal control. The data presented are the means ± SD from three independent experiments. * indicates P < 0.05, ** indicates P < 0.01, and **** indicates P < 0.0001, by unpaired t test.
Using raffinose as the bacteria's sole sugar source, we performed growth assays and quantitative real time RT-PCR assays. These assays demonstrated that blood isolates utilized raffinose better than their corresponding ear isolates (data not shown). We were curious if these differences impacted disease progression. With allelic exchange mutagenesis, I swapped the rafR alleles between the blood and ear isolates of serotype 14 ST15. These rafR swapped mutants showed a switch in their ability to utilize raffinose, suggesting that the SNP in rafR was driving the differential raffinose utilisation (Figure 2). Strikingly, in intranasal mouse challenge models, the rafR swapped strains also displayed a switch in their disease profiles. The rafR swapped blood isolate spread to the ear and brain, while rafR swapped ear isolate instead invaded the lungs (Figure 3). These results suggest that the ability to utilize raffinose plays a role in dictating tissue tropism and pneumococcal disease progression.
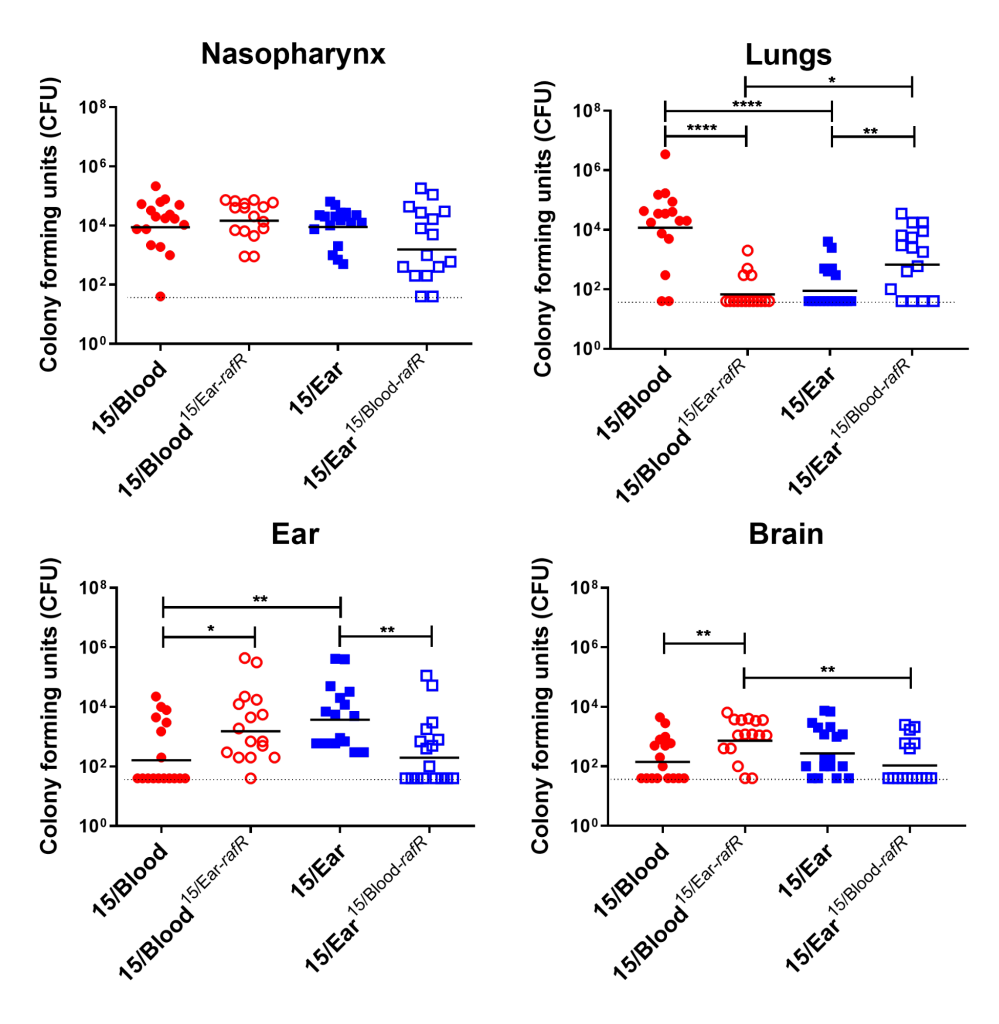
Figure 3. Virulence phenotype of Groups of 16 mice were infected intranasally with 108 CFU of the indicated strain. At 24 h, all mice from each group were euthanized and numbers of pneumococci in the indicated tissues/sites were quantified. Viable counts (total CFU per tissue) are shown for each mouse at each site; horizontal bars indicate the geometric mean (GM) CFU for each group; and the broken line indicates the threshold for detection. Differences in GM bacterial loads between groups are indicated by asterisks: * indicates P < 0.05, ** indicates P < 0.01, and **** indicates P < 0.0001, by unpaired t test.
Raffinose is a plant-derived trisaccharide sugar present in many staple foods, particularly beans and soy. Its influence on disease progression in this study is rather confounding, and the precise mechanism by which raffinose utilisation determines disease progression remains uncertain.10 Although humans are not readily able to metabolize it, dietary raffinose can be absorbed by the intestinal epithelium. This raises the possibility that at least small amounts of this sugar are present on mucosal surfaces.11 Another potential explanation for the results from this study is that rafR acts as a gene regulator. It's plausible that rafR acts on other downstream genes, in turn influencing the disease progression of these strains.
Role of Sugar Metabolism in Virulence
S. pneumoniae faces different environments within the human host, with each environment providing distinct carbohydrate sources.9 Thus, the ability to respond to and utilize different sugars must be crucial for both pneumococcal survival and disease progression. The importance of sugar metabolism is clear: the pneumococcus dedicates 30% of all transport mechanisms for carbohydrate import. It possesses at least 21 phosphotransferase systems (PTS) and up to 8 ABC transporters for this purpose.12 Other studies have shown that these carbohydrate transporters impact pneumococcal disease progression. For example, a sucrose PTS and an ABC transporter system of serotype 4 pneumococci each play roles in murine colonisation and pneumonia, respectively. In this same serotype, transporters for carbohydrates such as glucose, galactose and mannose impacted invasive pneumococcal disease.12 Yet, compared to other factors that influence pneumococcal virulence, carbohydrate metabolism is relatively understudied.
S. pneumoniae is a major cause of pneumonia and otitis media, as well as a cause of septicemia and meningitis, so this bacteria creates massive disease burdens. With the shortcomings of current vaccines and antibiotics, novel vaccines and therapeutics are urgently needed. Future research on the impact of sugar metabolism on the disease progression of S. pneumoniae will aid in this endeavour.
References
Editor's Note: The featured illustration for this post is credited to the CDC (Public Health Image Library, ID 22883). It was illustrated by Dan Higgins.
Vikrant Minhas is an academic who uses Benchling. Join him:
If you or your colleagues are interested in contributing an article about your research to the Benchling blog, let us know!