How AstraZeneca created an automated DNA assembly framework to support rapid and cost-efficient construct generation
Integration with Benchling unleashes the scale-up potential of a fully automated pipeline
While FRAGLER can and has been run offline, offline implementation doesn’t permit the type of scale that AstraZeneca needed. Although sequence fragmentation and fragment recycling can significantly drive down costs, that approach realizes its full potential only if you can find the fragments that you’re going to reuse. Öling explains, “There needs to be an algorithmic search to properly identify pre-existing fragments for FRAGLER to align. No human can keep track of everything if thousands of constructs are generated annually. Benchling allows us to achieve speed and scale by transforming previously manual processes relying on Excel and various in-house tools into fully automated steps.”
Leveraging Benchling's developer platform, the Benchling team built an integration to fully automate the process of generating de-novo fragments. It also allows for rapid searching of the Benchling database across thousands of pre-existing fragments in order to generate fully assembled in-silico constructs.
By integrating FRAGLER into Benchling, pre-existing fragments are identified and collected for the multiple sequence alignment and fragmentation steps. Benchling also creates pooling instructions for the resulting fragments that are used for construct assembly. Files containing information about the pools and which construct(s) they correspond to can then be entered back into the platform. In addition, Benchling facilitates QC steps, such as monitoring plasmid purity and concentration. The result for AstraZeneca is increased throughput coupled with high confidence in data quality since Benchling is the single source of truth for each step.
AstraZeneca developed an in-house design and request portal to enable global teams across Sweden and the U.K. to request constructs. Like FRAGLER, this portal was easily integrated with Benchling via the developer platform and open APIs. The entire process of DNA construct production, from initial request to sequence-validated plasmid, is automated and can be completed in about three weeks compared to 4-8 weeks (Figure 2). This is a timeline no DNA synthesis vendor can equal, says Öling, particularly for long, complex sequences such as SarsCoV or Cas9 expression constructs.
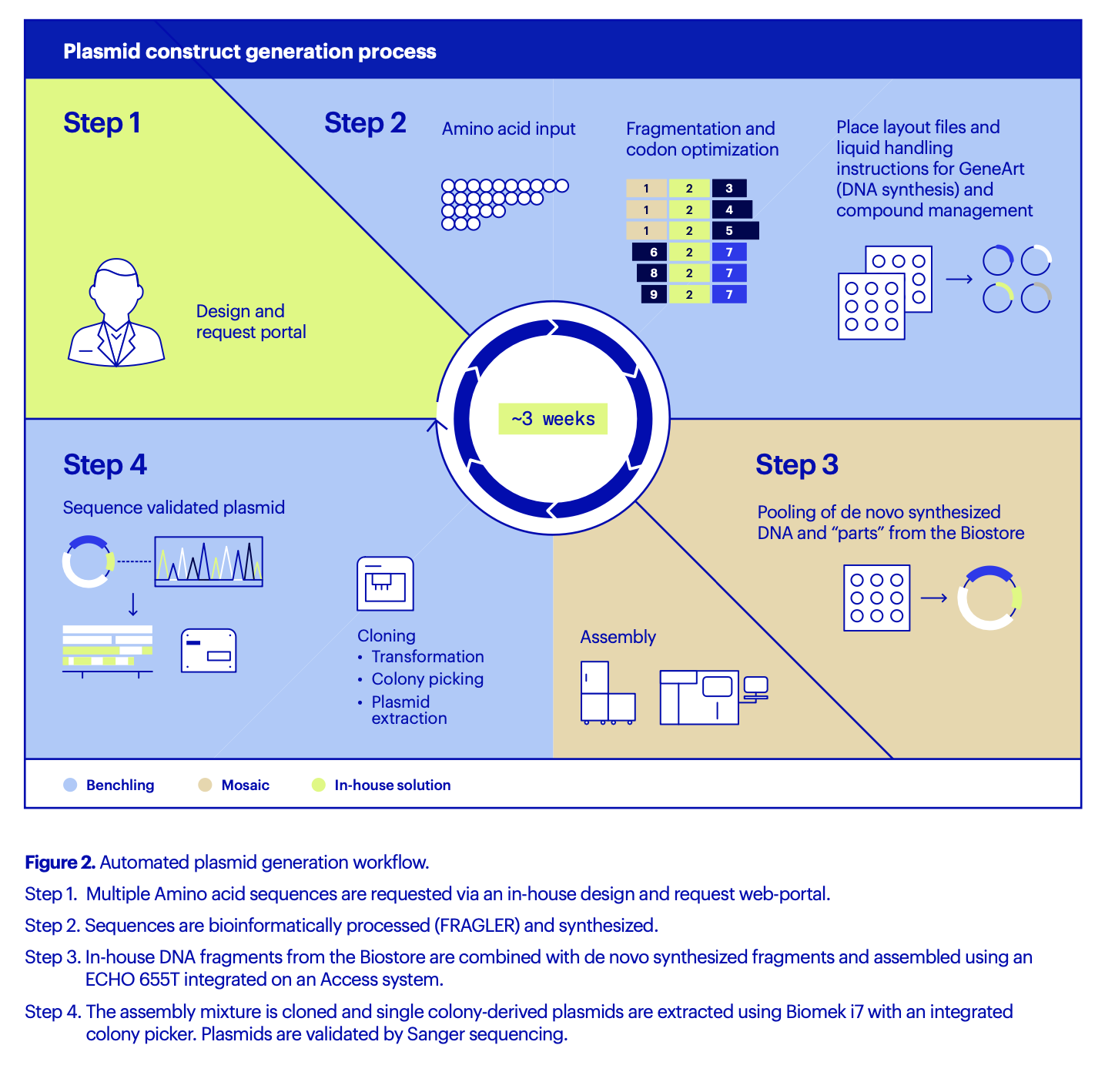
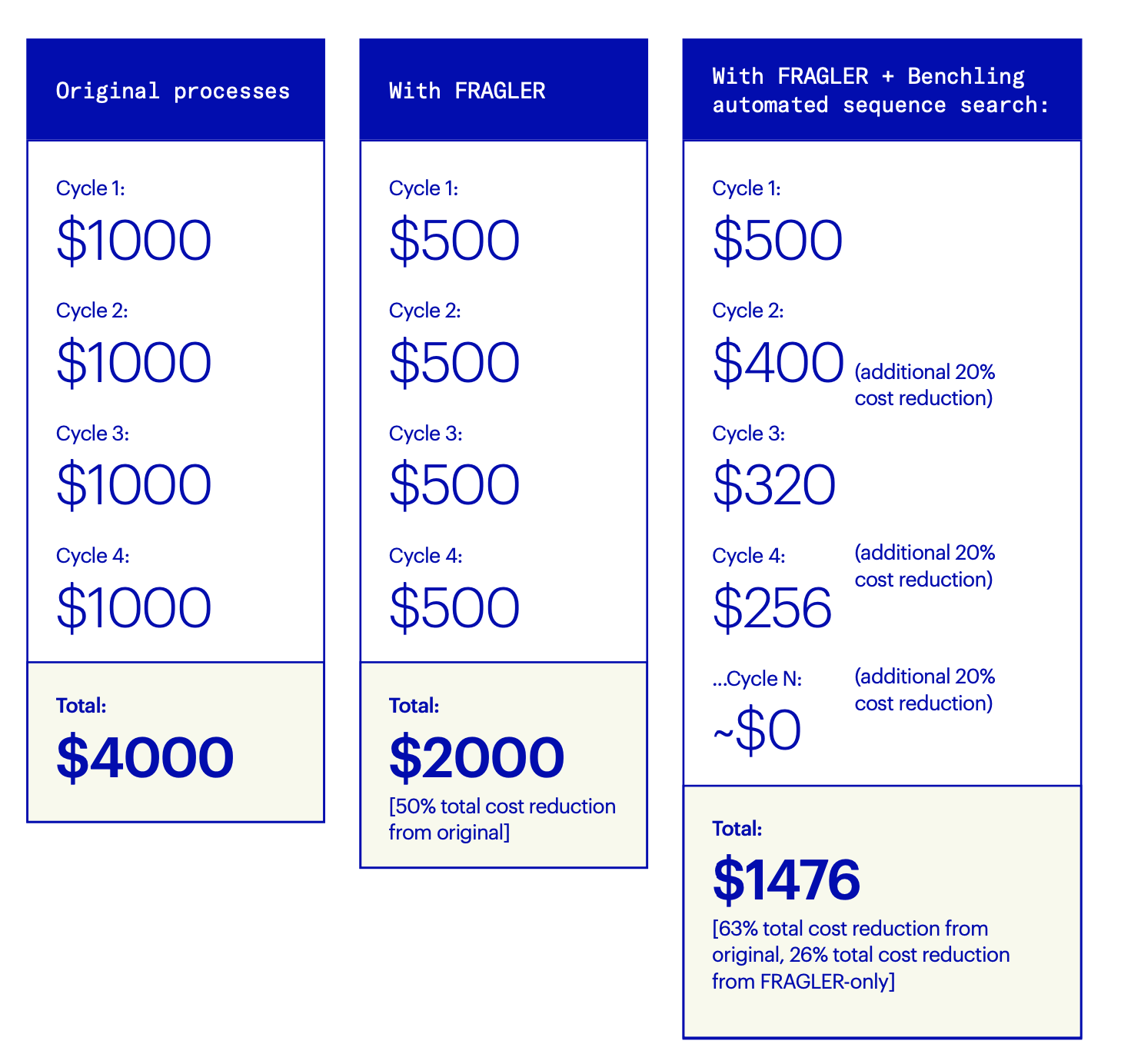
“The first time you run a campaign, FRAGLER will identify duplicate DNA sequences and help you save up to 50% of the cost of synthetic DNA synthesis by enabling fragment recycling,” says Öling. “But for each campaign after that,” he continues, “there is an estimated 10-30% additional cost reduction per iterative cycle, as more and more fragments are recycled and reused with each campaign. Eventually, most DNA fragments will be available in-house, meaning that you’ll reach nearly zero cost for generation of new synthetic DNA constructs. This, of course, can only be fully realized by the automated sequence search in Benchling, as manual searches for duplicate sequences at this scale are impossible.”
Powering breakthroughs for over 1,300 biotechnology companies, from startups to Fortune 500s
