Primer Design
Download "Primer Design" Ebook
Primers in RNA replication
It is the role RNA primers play in DNA synthesis that makes primer biology so incredibly important.In vivo, this is DNA replication or gene synthesis, a cornerstone of heredity where genetic material that was inherited from parent cells is copied for cell growth, division, and differentiation.
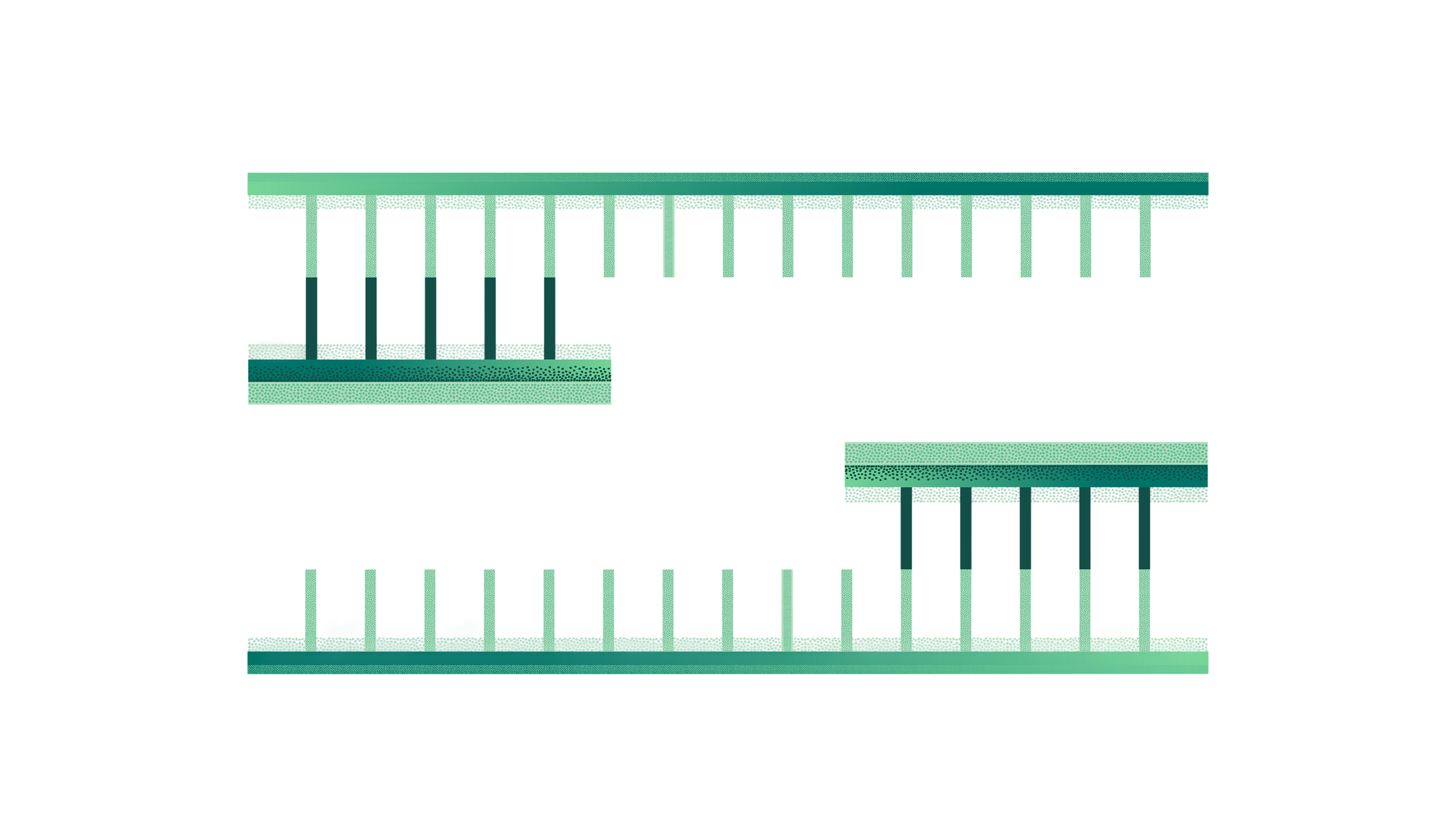
DNA replication begins at an origin of replication, which is identified by proteins called initiators that latch and make small openings in the double-stranded DNA double (dsDNA) helix. At these breaks, helicase enzymes break the bonds between the double strands, exposing single-stranded DNA (ssDNA) in a Y-shaped end called the DNA replication fork. The replication fork is where DNA replication will actually occur, providing two strands of parent DNA now exposed to act as the DNA template (gray strands below). Next, RNA primase lays down the RNA primer on each of the parent template strands, providing the 3’ end for a new polynucleotide strand to begin forming. DNA polymerase binds and synthesis is initiated, with daughter strands being formed (purple strands). In this diagram, the top daughter strand is called the leading strand and is exposed in the 5 prime (5’) → 3’ prime (3’) direction right to left, allowing DNA synthesis to occur continuously as the helix unwinds.
Lagging strand synthesis of the bottom daughter strand occurs in the 5’ → 3’ direction left to right, where DNA synthesis must continually start over as the helix unwinds. This creates many short fragments of synthesized products, known as Okazaki fragments which average ~150-200 nucleotides long. Each Okazaki fragment contains an RNA-DNA joint as a consequence of RNA priming, a discovery that was important evidence for determining the role of RNA primers in DNA replication. Okazaki fragments must be joined together by the enzyme ligase, which catalyzes the formation of covalent phosphodiester bonds between two strands of DNA, ligating them together.
Forward and Reverse Primers
Primers only bind to single-stranded DNA. In DNA replication in vivo, this is made available at the replication fork. In vitro, scientists can create ssDNA by denaturing dsDNA with heat, effectively breaking hydrogen bonds that hold the two strands together. This creates two linear template strands where DNA synthesis can occur continuously on both strands, often called the top and bottom strand in place of the terms leading and lagging strands.
A pair of primers must be used, one for the top strand and one for the bottom. The primer pair binds on opposite ends of the sequence being amplified, with their 3’ ends pointing toward each other. The oligonucleotide primers being designed do not actually have to hybridize completely to the template strand; it is essential, however, that the 3’ end corresponds completely to the template DNA for elongation to occur. The forward primer anneals to the bottom strand that runs 3’ to 5’, and the reverse primer anneals to the complementary top strand of DNA that runs 5’ to 3’. After elongation, this results in two new strands of double-stranded DNA: one made from the forward primer; and the other made from the reverse primer.
In order for the primers to bind to the template strand, it’s important they are not complementary to each other. Primers complementary to each other can result in the creation of primer dimers, which are two primers binding to each other instead of the template strand. Primer dimers are discussed in detail in the section on what to avoid when designing primers for PCR.
Powering breakthroughs for over 1,300 biotechnology companies, from startups to Fortune 500s
